Articles
- Page Path
- HOME > Osong Public Health Res Perspect > Volume 7(6); 2016 > Article
-
Original Article
Rapid Molecular Approach for Simultaneous Detection ofSalmonella spp.,Shigella spp., andVibrio cholera - Reza Ranjbara, Ali Naghonia, Davoud Afsharb, Farhad Nikkhahic, Mohsen Mohammadid
-
Osong Public Health and Research Perspectives 2016;7(6):373-377.
DOI: https://doi.org/10.1016/j.phrp.2016.10.002
Published online: November 5, 2016
aMolecular Biology Research Center, Baqiyatallah University of Medical Sciences, Tehran, Iran
bDepartment of Microbiology, Faculty of Medicine, Zanjan University of Medical Sciences, Zanjan, Iran
cDepartment of Pathobiology, School of Public Health, Tehran University of Medical Sciences, Tehran, Iran
dDepartment of Pharmaceutical Biotechnology, Faculty of Pharmacy, Lorestan University of Medical Sciences, Khorramabad, Iran
- ∗Corresponding author. ranjbarre@gmail.com
• Received: August 21, 2016 • Revised: September 22, 2016 • Accepted: October 10, 2016
Copyright © 2016 Korea Centers for Disease Control and Prevention. Published by Elsevier Korea LLC.
This is an open access article under the CC BY-NC-ND license (http://creativecommons.org/licenses/by-nc-nd/4.0/).
Abstract
-
Objectives
- Gastrointestinal tract infection is still one of the serious public health problems in many geographic areas and is endemic in most countries including Iran. Early detection of the gastrointestinal tract pathogens can be extremely important. The aim of the current study was to apply a shortened time-multiplex polymerase chain reaction (PCR) for rapid and simultaneous detection of Salmonella spp., Shigella spp., and Vibrio cholera.
-
Methods
- The standard and clinical strains of Salmonella spp., Shigella spp., and V. cholerae were used in the assay. Multiplex PCR was performed and optimized based on amplification of invA, putative integrase, and ompW genes for detecting Salmonella spp., Shigella spp., and V. cholerae, respectively. The specificity of the assay was evaluated by testing 12 different bacterial species.
-
Results
- Only Salmonella spp., Shigella spp., and V. cholerae strains had positive results when subjected to the assay using multiplex PCR. The assay showed a high sensitivity, and no amplification products were observed in multiplex PCR with any of the other microorganisms.
-
Conclusion
- Our study indicated that the invA, putative integrase, and ompW-based multiplex PCR assay appears to be an efficient method for rapid and simultaneous detection of Salmonella spp., Shigella spp., and V. cholerae.
- Worldwide, gastrointestinal tract infections are the second most important cause of death; about 25 million enteric infections occur each year. These infections cause significant morbidity and death in children younger than 5 years in particular and in elderly people. It has been estimated that 4–6 million children die each year because of diarrheal diseases, particularly in the developing countries [1]. Numerous outbreaks of diarrheal illness caused by various microorganisms have been reported. Microorganisms such as Shigella, Salmonella, Vibrio, Escherichia coli, Campylobacter jejuni, Giardia lamblia, Cryptosporidium, and Rotaviruses have been reported to be the most important causes of diarrheal outbreaks. Salmonella spp., Shigella spp., and V. cholerae are the most important bacterial causes of diarrhea in Iran 2, 3, 4, 5.
- The diseases caused by all of these microorganisms could be serious, resulting in death. V. cholerae causes cholera, a disease with endemic or pandemic potential characterized by watery diarrhea and vomiting, leading to severe and rapidly progressing dehydration and shock [6]. The symptoms are caused by cholera toxin, which is produced by pathogenic strains of V. cholerae. Many efforts have been made to introduce a more effective vaccine, but many researches have shown that the vaccination has no role for cholera; however, new oral vaccines are displaying egregious promise [7].
- Shigellosis and salmonellosis are caused by Shigella spp. and Salmonella spp., respectively. These organisms are likely to be the common cause of diarrhea worldwide. Shigella spp. are the causative agents of inflammatory diarrhea and dysentery, thus presenting a serious challenge to public health authorities worldwide [5]. Although shigellosis has no known animal reservoirs, we are still lacking an effective vaccine owing to poor immune responses to oral vaccines and existence of multiple serotypes [8].
- Unlike Shigella, Salmonella spp. (except Salmonella enterica subspecies Typhi) are found in many animals. Thus, salmonellosis is well recognized as zoonosis disease [9]. The prevalence of Salmonella infection varies depending on the waste disposal, water supply, food preparation practices, and climate. Gastroenteritis is the most common disease among children caused by Salmonella [5].
- The traditional methods for detection of bacterial infections are still primarily based on culture and serological methods that may take several days to be completed. There has been a general move toward molecular methods for microbial detection, which are based less on phenotypic features and more on stable genotypic characteristics. In recent years, polymerase chain reaction (PCR) and similar nucleotide-based methods have become potentially powerful alternative approaches in microbiological diagnostics because of their higher user-friendliness, rapidity, reproducibility, accuracy, and affordability. These methods have also gained momentum in terms of use for rapid, specific, and sensitive detection of foodborne pathogens 10, 11, 12, 13, 14, 15.
- Multiplex polymerase chain reaction is a variant of PCR in which two or more loci are simultaneously amplified in the same reaction [16]. This technique is a powerful molecular method in microbiological diagnostics that allows the simultaneous amplification of more than one target sequence in a single PCR reaction, saving considerable time and effort, and decreasing the number of reactions to be performed in order to assess the possible presence of foodborne pathogens 16, 17, 18.
- In this study, we describe a multiplex PCR assay for the rapid and simultaneous detection of Salmonella spp., Shigella spp., and Vibrio cholera.
Introduction
- 2.1 Bacterial strains
- The bacterial strains were obtained from the Pasteur Institute, Tehran, Iran and used in this study (Table 1). Clinical isolates of the three most important foodborne bacterial pathogens including Salmonella and Shigella were obtained from patients admitted to Children's Medical Center and Baqiyatallah Hospitals in Tehran, Iran, during 2012–2014. Subsequently, identification of the references and clinical strains was confirmed by culture, biochemical testing by the API test system (BioMérieux, Marcy-l'Étoile, France), and slide agglutination with serovar specific antisera (Staten Serum Institute, Copenhagen, Denmark). V. cholerae isolates were provided by the Molecular Biology Research Center affiliated to Baqiyatallah Hospital.
- All bacterial strains were grown either on Brain Heart Infusion (BHI; Difco Laboratories, Detroit, MI, USA) or Luria–Bertani (LB) broth (Merck, Darmstadt, Germany) at 37°C for 18–24 hours.
- 2.2 DNA extraction
- Genomic DNAs from all microorganisms were extracted using the DNA extraction kit (DNP, DNA Extraction Kit; Cinagene Company, Tehran, Iran) according to the manufacturer’s instructions. DNA concentration and purity were spectrophotometrically assessed by reading A260 and A280 and confirmed by visualization on 1% agarose gel. Then, DNA was diluted to 1 mg/mL in nuclease-free water and stored at –20°C until required for analysis.
- 2.3 Primers and multiplex PCR conditions
- The AlleleID software version 7.01 (Premier Biosoft Int., Palo Alto, CA, USA) was used for all oligonucleotide primers designed in this study. All primers were purchased from Bioneer (Daejeon, South Korea). The in silico specificity was analyzed using the Basic Local Alignment Search Tool (BLAST) from the GenBank database. The characteristics of the primers used for multiplex PCR are given in Table 2.
- PCR was carried out with a 50-μL mixture containing 10mM Tris–HCl (pH 8.3), 50mM.
- In this study, we used KCl, 1.5mM MgCl2, 1 U of Taq DNA polymerase (Promega, Madison, WI, USA), 0.2mM deoxynucleoside triphosphate, a 0.1μM concentration of each primers, and 5 μL of the DNA sample.
- Multiplex PCR was performed under the following conditions: 35 cycles with heat denaturation at 95°C for 30 seconds, primer annealing at 60°C for 30 seconds, and DNA extension at 72°C for 60 seconds in Eppendorf gradient master cycler (Roche, Mannheim, Germany). The amplified DNA was separated by 1% agarose gel electrophoresis, stained with ethidium bromide, and visualized by UV transilluminator.
- 2.4 Sensitivity and specificity
- To determine the sensitivity of the multiplex PCR assay, 10-fold serial dilutions were made from extracted genomic DNA (498 ng/μL), and the detection limit of the multiplex PCR was defined as the lowest concentration of DNA that could be amplified. The specificity of multiplex PCR was evaluated using three species including Staphylococcus aureus PTCC (Persian Type Culture Collection) 1189, E. coli ATCC (American Type Culture Collection) 25922, and Citrobacter freundii ATCC 8090 as negative controls.
Materials and methods
- The multiplex PCR using three sets of primer pairs targeted for the invA, putative integrase, and ompW genes, correctly identified Salmonella spp., Shigella spp., and V. cholerae and differentiated them by the different-size bands products: three positive bands, which consist of invA (403 bp), putative integrase (159 bp), and ompW (592 bp) PCR products (Figure 1). No amplification products were observed in multiplex PCR with any of the other microorganisms subjected to the assay (Table 1). The sensitivity of the multiplex PCR was assessed to be 5 ng/μL of the pure DNA.
Results
- Salmonella spp., Shigella spp., and V. cholerae are responsible for large numbers of intestinal infections in humans worldwide. Molecular techniques, such as multiplex PCR, are proving useful in detection of pathogens in a wide spectrum of matrices 10, 19. This technique enables us to identify these three pathogens at one experiment, obviating the need for three separate experiments. The use of multiplex PCR substantially reduces the time and manpower required when compared with conventional methodologies. Here, we report a multiplex PCR assay for detection of Salmonella, Shigella, and V. cholerae based on invA, ompW, and putative integrase genes, respectively. Previous studies indicated that these genes are conserved in each species. Many studies noted that invA is a specific and sensitive target for detection of Salmonella spp. 20, 21. Also, the ompW gene has been previously used for identification of V. cholerae, owing to its specificity [22]. Furthermore, restriction fragment length polymorphism analysis and nucleotide sequence data have shown that the ompW gene is highly conserved among all V. cholerae biotypes, suggesting the ompW gene can be considered a good target for the specific identification of V. cholerae strains [23].
- Unlike the above-mentioned species, Shigella genomes have a high level of similarity with the E. coli genome; hence, the whole sequences of Shigella dysenteriae, Shigella boydii, Shigella flexneri, and Shigella sonnei have ∼3 Mb of genomic DNA in common with all sequenced E. coli genomes [24]. However, based on the comparative genomic analysis, a specific target known as putative integrase locus, conserved in all Shigella species, was subjected to identification of Shigella species. Hence, Shigella-specific primers were designed based on putative integrase locus. The results also showed that this locus is a suitable target for specific identification of Shigella species.
- In many research studies, multiplex PCR has been applied for rapid identification of diarrheal agents 25, 26. All of these studies noted that multiplex PCR is a reliable, useful, and cost-effective assay, which is consistent with our results. Jin et al [27] studied foodborne pathogenic bacteria including C. jejuni, Shigella, Salmonella, Vibrio parahaemolyticus, S. aureus, E. coli O157:H7, and several other bacterial species and showed that multiplex PCR is time-saving assay in comparison with conventional PCR. Furthermore, Paniagua et al [28], who described the detection of different foodborne pathogens by multiplex PCR, noted that this method could be useful for quick detection of foodborne pathogens.
- There are inconsistent reports about the sensitivity of multiplex PCR. According to Tsai et al [29], the sensitivity of multiplex PCR is considerably lower than that of monoplex PCR because of the primers’ interference, so that it can be decreased several times compared with conventional PCR. However, Al-Talib et al [30] showed that multiplex PCR has a high level of sensitivity, and it might be useful as an alternative diagnostic tool for diarrheal diseases. In our study, a high level of sensitivity (5 ng/μL) was also observed. It appears that the sensitivity of multiplex PCR is related to primer length and can be enhanced by shortening the primers’ length. However, this modification leads to low specificity.
- The infections caused by enteric pathogens comprise second commonest medical problems after respiratory infectious disease 31, 32. Salmonella, Shigella, and Vibrio are among the most prevalent and endemic food and water- borne pathogens in Iran 33, 34, 35, 36. Rapid and simultaneous detection of these common bacteria in is extremely important to ensure food and water safety. For this purpose, we developed and successfully applied a multiplex PCR for the rapid identification of Salmonella spp., Shigella spp., and V. cholera. This technique decreases the test time of PCR. This method is simple and rapid, and the results obtained proved to be highly specific and sensitive and can be expanded to additional species. Moreover, multiplex PCR may provide an epidemiological tool to investigate the wide spread of diarrheagenic pathogens in various areas worldwide.
Discussion
- The authors declare that there is no conflict of interests.
Conflicts of interest
- 1. Guerrant R.L., Hughes J.M., Lima N.L.. Diarrhea in developed and developing countries: magnitude, special settings, and etiologies. Rev Infect Dis 12:1990;S41−S50. PMID: 2406855.ArticlePubMedPMC
- 2. Ranjbar R., Afshar D., Mehrabi Tavana A.. Development of multiplex PCR for simultaneous detection of three pathogenic Shigella species. Iran J Public Health 43:2014;1657−1663. PMID: 26171358.PubMedPMC
- 3. Ranjbar R., Rahbar M., Naghoni A.. A cholera outbreak associated with drinking contaminated well water. Arch Iran Med 14:2011;339−340. PMID: 21888459.PubMed
- 4. Ranjbar R., Naghoni A., Farshad S.. Use of TaqMan® real-time PCR for rapid detection of Salmonella serotypes Typhi and Typhimurium. Acta Microbiol Immunol Hung 61:2014;121−130. PMID: 24939681.ArticlePubMed
- 5. Mulatu G., Beyene G., Zeynudin A.. Prevalence of Shigella, Salmonella and Campylobacter species and their susceptibility patters among under five children with diarrhea in Hawassa town, south Ethiopia. Ethiop J Health Sci 24:2014;101−108. PMID: 24795510.ArticlePubMedPMC
- 6. Akoachere J.F.T., Mbuntcha C.K.. Water sources as reservoirs of Vibrio cholerae O1 and non-O1 strains in Bepanda, Douala (Cameroon): relationship between isolation and physico-chemical factors. BMC Infect Dis 14:2014;421PMID: 25073409.ArticlePubMedPMC
- 7. Sack D.A., Sack R.B., Nair G.B.. Cholera. Lancet 17:2004;223−233.Article
- 8. Kweon M.N.. Shigellosis: the current status of vaccine development. Curr Opin Infect Dis 2:2008;313−318.Article
- 9. Okoro C.K., Kingsley R.A., Connor T.R.. Intra-continental spread of human invasive Salmonella Typhimurium pathovariants in sub-Saharan Africa. Nat Genet 44:2012;1215−1221. PMID: 23023330.ArticlePubMedPMC
- 10. Karami A., Ranjbar R., Ahmadi Z.. Rapid detection of different serovares of Salmonella entrica by Multiplex PCR. Iran J Publ Health 36:2007;38−42.
- 11. Malorny B., Hoorfar J., Bunge C.. Multicenter validation of the analytical accuracy of Salmonella PCR: towards an international standard. Am Soc Microbiol 69:2003;290−296.ArticlePDF
- 12. Popovic T., Bopp C., Olsvik O.. Epidemiologic application of a standardized ribotype scheme for Vibrio cholerae O1. J Clin Microbiol 31:1993;2474−2482. PMID: 7691876.ArticlePubMedPMC
- 13. Vargas M., Gascon J., Jimenez De Anta M.T.. Prevalence of Shigella enterotoxins 1 and 2 among Shigella strains isolated from patients with traveler's diarrhea. J Clin Microbiol 37:1999;3608−3611. PMID: 10523561.ArticlePubMedPMC
- 14. Roy S., Thanasekaran K., Dutta Roy A.R.. Distribution of Shigella enterotoxin genes and secreted autotransporter toxin gene among diverse species and serotypes of Shigella isolated from Andaman Islands, India. Trop Med Int Health 11:2006;1694−1698. PMID: 17054749.ArticlePubMed
- 15. Orrett F.A.. Prevalence of Shigella serogroups and their antimicrobial resistance patterns in southern Trinidad. J Health Popul Nutr 26:2008;456−462. PMID: 19069625.PubMedPMC
- 16. Akiba M., Kusumoto M., Iwata T.. Rapid identification of Salmonella enterica serovars, Typhimurium, Choleraesuis, Infantis, Hadar, Enteritidis, Dublin and Gallinarum, by multiplex PCR. J Microbiol Methods 85:2011;9−15. PMID: 21329737.ArticlePubMed
- 17. Kong R.Y., Lee S.K., Law T.W.. Rapid detection of six types of bacterial pathogens in marine waters by multiplex PCR. Water Res 36:2002;2802−2812. PMID: 12146868.ArticlePubMed
- 18. Amini K., Zahraei-Salehi T., Nikbakht G.. Molecular detection of invA and spv virulence genes in Salmonella enteritidis isolated from human and animals in Iran. Afr J Microbiol Res 4:2010;2202−2210.
- 19. Bohaychuk V.M., Gensler G.E., McFall M.E.. A real-time PCR assay for the detection of Salmonella in a wide variety of food and food–animal matrices. J Food Prot 70:2007;1080−1087. PMID: 17536664.ArticlePubMed
- 20. Rahn K., De Grandis S.A., Clarke R.C.. Amplification of an invA gene sequence of Salmonella typhimurium by polymerase chain reaction as a specific method of detection of Salmonella. Mol Cell Probes 6:1992;271−279. PMID: 1528198.ArticlePubMed
- 21. Mainar-Jaimea R.C., Andrésa S., Vicoa J.P.. Sensitivity of the ISO 6579:2002/Amd 1:2007 standard method for detection of Salmonella spp. on mesenteric lymph nodes from slaughter pigs. J Clin Microbiol 51:2013;89−94. PMID: 23100334.ArticlePubMedPMC
- 22. Wei S., Zhao H., Xian Y.. Multiplex PCR assays for the detection of Vibrio alginolyticus, Vibrio parahaemolyticus, Vibrio vulnificus, and Vibrio cholerae with an internal amplification control. Diagn Microbiol Infect Dis 79:2014;115−118. PMID: 24731836.ArticlePubMed
- 23. Nandi B., Nandy R.K., Mukhopadhyay S.. Rapid method for species-specific identification of Vibrio cholerae using primers targeted to the gene of outer membrane protein OmpW. J Clin Microbiol 38:2000;4145−4151. PMID: 11060082.ArticlePubMedPMC
- 24. Yang F., Yang J., Zhang X.. Genome dynamics and diversity of Shigella species, the etiologic agents of bacillary dysentery. Nucleic Acids Res 33:2005;6445−6458. PMID: 16275786.ArticlePubMedPMC
- 25. Chen J., Tang J., Liu J.. Development and evaluation of a multiplex PCR for simultaneous detection of five foodborne pathogens. J Appl Microbiol 112:2012;823−830. PMID: 22268759.ArticlePubMed
- 26. Vondrakova L., Pazlarova J., Demnerova K.. Detection, identification and quantification of Campylobacter jejuni, coli and lari in food matrices all at once using multiplex qPCR. Gut Pathog 6:2014;12PMID: 25057300.ArticlePubMedPMC
- 27. Jin S.Q., Yin B.C., Ye B.C.. Multiplexed bead-based mesofluidic system for detection of food-borne pathogenic bacteria. Am Soc Microbiol 75:2009;6647−6654.Article
- 28. Paniagua G.L., Monroy E., García-González O.. Two or more enteropathogens are associated with diarrhoea in Mexican children. Ann Clin Microbiol Antimicrob 6:2007;17PMID: 18162140.ArticlePubMedPMC
- 29. Tsai Y.L., Tran B., Sangermano L.R.. Detection of poliovirus, hepatitis A virus, and rotavirus from sewage and ocean water by triplex reverse transcriptase PCR. Appl Environ Microbiol 60:1994;2400−2407. PMID: 8074520.ArticlePubMedPMC
- 30. Al-Talib H., Latif B., Mohd-Zain Z.. Pentaplex PCR assay for detection of hemorrhagic bacteria from stool samples. J Clin Microbiol 52:2014;3244−3249. PMID: 24958797.ArticlePubMedPMC
- 31. Ranjbar R., Soltan Dallal M.M., Pourshafie M.R.. Serogroup distribution of Shigella spp. in Tehran. Iranian J Publ Health 33:2004;32−35.
- 32. Ranjbar R., Salimkhani E., Sadeghifard N.. An outbreak of gastroenteritis of unknown origin in Tehran. Pak J Biol Sci 10:2007;1138−1140. PMID: 19070067.ArticlePubMed
- 33. Ranjbar R., Rahbar M., Naghoni A.. A cholera outbreak associated with drinking contaminated well water. Arch Iran Med 14:2011;339−340. PMID: 21888459.PubMed
- 34. Pourshafie M.R., Bakhshi B., Ranjbar R.. Dissemination of a single Vibrio cholerae clone in cholera outbreaks during 2005 in Iran. J Med Microbiol 56:2007;1615−1619. PMID: 18033829.ArticlePubMed
- 35. Ranjbar R., Giammanco G.M., Farshad S.. Serotypes, antibiotic resistance, and class 1 integrons in Salmonella isolates from pediatric cases of enteritis in Tehran, Iran. Foodborne Pathog Dis 8:2011;47−53.Article
- 36. Ranjbar R., Hosseini M.J., Kaffashian A.R.. An outbreak of shigellosis due to Shigella flexneri serotype 3a in a prison in Iran. Arch Iran Med 13:2010;413−416. PMID: 20804308.PubMed
References
Figure 1The multiplex PCR results. Lanes 2–4, uniplex PCR of some representative strains of Vibrio cholera, Salmonella spp., and Shigella spp., respectively. Lane 1, multiplex PCR for the same three bacterial strains in a single PCR tube. M = molecular weight (100 bp DNA ladder); PCR = polymerase chain reaction.
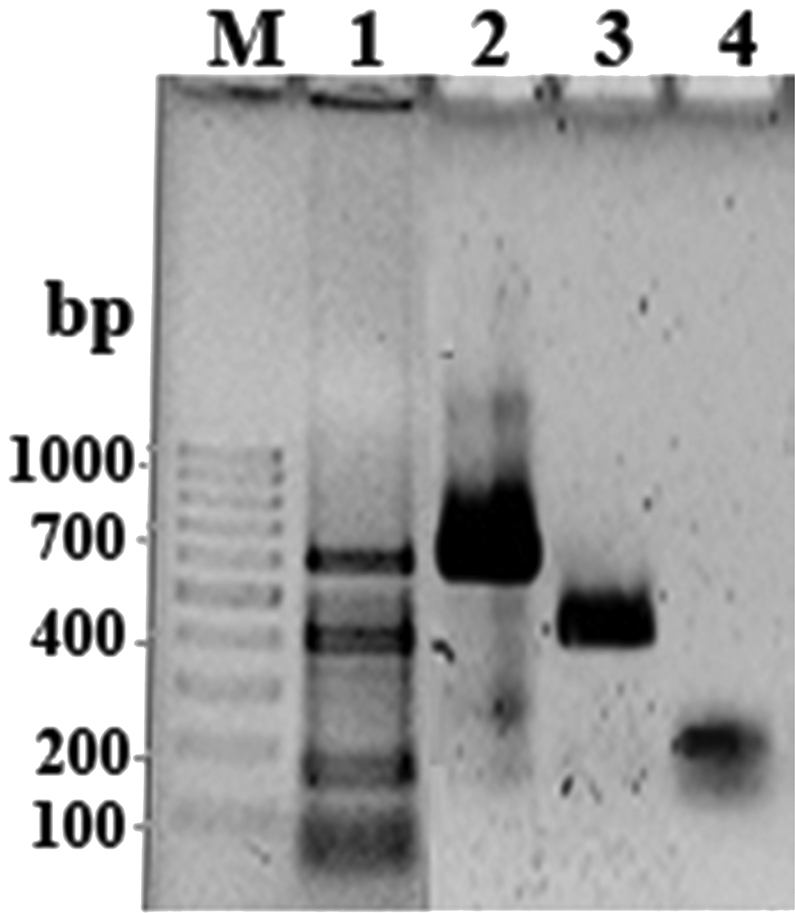

Table 1Bacterial strains included in this study, and performance of the multiplex PCR assay for detecting Salmonella, Shigella, and Vibrio cholera.
Table 2Primers sequences used for amplification by multiplex PCR.
| Primer name | Sequence (5′→3′) | Product (bp) | Target | Reference |
|---|---|---|---|---|
|
Vibrio-F Vibrio-R |
ATAATGGCTCACCAAGAAGG TTAGAACTTATAACCACC |
592 | ompW | This study |
|
Shigella-F Shigella -R |
TCCGTCATGCTGGATGAACGATGT ACAGTTCAGGATTGCCCGAGACACA |
159 | Putative integrase | Ranjbar et al [2] |
|
Salmonella-F Salmonella-R |
GTATTGTTGATTAATGACATCCG ATATTACGCTACGGAAACACGTT |
403 | invA | This study |
Figure & Data
References
Citations
Citations to this article as recorded by 

- Development and Validation of an Efficient Multiplex PCR Assay for Simultaneous Detection of Six Common Foodborne Pathogens and Hygiene Indicators
Natharin Ngamwongsatit, Soraya Chaturongakul, Ratchaneewan Aunpad
Foodborne Pathogens and Disease.2023; 20(6): 222. CrossRef - Rapid and multiplexed quantification of Salmonella, Escherichia coli O157:H7, and Shigella flexneri in ground beef using flow cytometry
Ziquan Wang, Qian Xu, Siyuan Liu, Yingying Liu, Ying Gao, Meng Wang, Ling Zhang, Haiyan Chang, Qiang Wei, Zhiwei Sui
Talanta.2022; 238: 123005. CrossRef - Development of multiplex real-time quantitative PCR for simultaneous detection of Chlamydia trachomatis, Mycoplasma hominis, Ureaplasma urealyticum, and Mycoplasma genitalium in infertile women
Sara Sadeqi, Farhad Nikkhahi, Amir Javadi, Sonia Eskandarion, Seyed Mahmoud Amin Marashi
Indian Journal of Medical Microbiology.2022; 40(2): 231. CrossRef - Multiple fluorescent saltatory rolling circle amplification (SRCA) for simultaneous and sensitive detection of Salmonella spp. and Shigella spp. in food
Wei Guo, Qian Yang, Jie Liu, Xiuling Chen, Yunzhe Zhang, Wei Zhang
LWT.2022; 168: 113875. CrossRef - Campylobacter Species in the Middle East
Daryoush Babazadeh, Reza Ranjbar
Journal of Veterinary Physiology and Pathology.2022; 1(1): 1. CrossRef - Development of rapid gold nanoparticles based lateral flow assays for simultaneous detection of Shigella and Salmonella genera
Mohammad Lukman Yahaya, Nor Dyana Zakaria, Rahmah Noordin, Khairunisak Abdul Razak
Biotechnology and Applied Biochemistry.2021; 68(5): 1095. CrossRef - Ultrasensitive pathogen detection with a rolling circle amplification-empowered multiplex electrochemical DNA sensor
Cheryl S.Y. Yeap, Thanyarat Chaibun, Su Yin Lee, Bin Zhao, Yuan Jan, Chan La-o-vorakiat, Werasak Surareungchai, Shiping Song, Benchaporn Lertanantawong
Chemical Communications.2021; 57(91): 12155. CrossRef - Taqman hydrolysis probe application for Escherichia coli, Salmonella enterica, and Vibrio cholerae detection in surface and drinking water
Ahmed K. A. El-Sayed, Mohamed I. Abou-Dobara, Camelia A. Abdel-Malak, Amira A. E. El-Badaly
Journal of Water, Sanitation and Hygiene for Devel.2019; 9(3): 492. CrossRef - Pathways of healthcare and antibiotics use following reported gastrointestinal illness: a cross-sectional study in rural Anhui, China
Xing Rong Shen, Maomao Xie, Jing Chai, Rui Feng, Jing Cheng, Rong Liu, Paul Kadetz, DeBin Wang
BMJ Open.2019; 9(8): e030986. CrossRef - DNA Microarray for Rapid Detection and Identification of Food and Water Borne Bacteria: From Dry to Wet Lab
Reza Ranjbar, Payam Behzadi, Ali Najafi, Raheleh Roudi
The Open Microbiology Journal.2017; 11(1): 330. CrossRef


 PubReader
PubReader Cite
Cite
