Articles
- Page Path
- HOME > Osong Public Health Res Perspect > Volume 8(1); 2017 > Article
-
Brief Report
Emergence of Norovirus GII.17-associated Outbreak and Sporadic Cases in Korea from 2014 to 2015 - Sunyoung Jung, Bo-Mi Hwang, HyunJu Jung, GyungTae Chung, Cheon-Kwon Yoo, Deog-Yong Lee
-
Osong Public Health and Research Perspectives 2017;8(1):86-90.
DOI: https://doi.org/10.24171/j.phrp.2017.8.1.12
Published online: February 28, 2017
Division of Enteric Diseases, Center for Infectious Diseases, National Research Institute of Health, Osong, Korea
- Corresponding author: Deog-Yong Lee, E-mail: leedy0610@korea.kr
Copyright © 2017 Korea Centers for Disease Control and Prevention
This is an open access article under the CC BY-NC-ND license (http://creativecommons.org/licenses/by-nc-nd/4.0/).
Abstract
- Human norovirus are major causative agent of nonbacterial acute gastroenteritis. In general, genogroup (G) II.4 is the most prominent major genotype that circulate in human population and the environment. However, a shift in genotypic trends was observed in Korea in December 2014. In this study, we investigated the trend of norovirus genotype in detail using the database of Acute Diarrhea Laboratory Surveillance (K-EnterNet) in Korea. GII.17 has since become a major contributor to outbreaks of norovirus-related infections and sporadic cases in Korea, although the reason for this shift remain unknown.
- Norovirus is a major causative agent of acute viral gastroenteritis in humans of all ages during the winter season. Noroviruses can be classified into 5 genogroups (GI–GV) based on genetic diversity, of the polymerases and capsid genes. GI, II, and IV are known to infect humans, and GII.4, which undergoes rapid antigenic variation, is the most predominant genotype worldwide [1]. This genotype has also been reported to cause both sporadic cases and outbreaks and was detected in humans and the environment in Korea [2,3]. However, several recent outbreaks have been attributed to GII.17, and the prevalence of this genotype in both outbreaks and sporadic cases has increased sharply in Korea [4]. Therefore, we hereby report trends in norovirus genotypes in Korea.
INTRODUCTION
- We analyzed the proportions of norovirus genotypes detected in Korea from 2012 to 2015. Genotype data were collected from outbreaks and sporadic cases. An outbreak was defined that patients were occurred more than two people by norovirus infection at the same time and place [5]. Fecal samples were collected from diarrheal patients by epidemiologists in nationwide. Sporadic cases were used for sentinel acute diarrheal laboratory surveillance, a service in which epidemic pathogens were monitored using diarrheal fecal samples collected from 72 collaborating hospitals. All specimens (outbreak and sporadic) were tested for norovirus infection at 17 regional Institutes of Health and Environment Research. Partial capsid nucleotides were analyzed at sequencing centers and automatically registered in a web reporting system. K-EnterNet is a web-based reporting system and sequence analysis tool operated by the Korea National Institute of Health. This system collected and separately recorded epidemic and partial capsid nucleotide sequence data from sporadic and outbreak cases. To investigate the prevalence of each norovirus genotype, we extracted outbreak (n = 1,941) and sporadic (n = 3,931) nucleotide sequences from K-EnterNet from the past 42-month period (from 2012 to 2015) in Korea.
- Primary norovirus infection screening was performed via multiplex real-time polymerase chain reaction (RT-PCR; Ac-cuPower® Norovirus RT-PCR kit, Bioneer, Daejeon, Korea). Genotyping was performed using K-EnterNet and confirmed using the NoroNet typing tool (http://www.rivm.nl) after semi-nested RT-PCR (AccuPower® HotStart Pre Mix; Bioneer) and automated sequencing (ABI3730XL; Applied Biosystems, Foster City, CA, USA). Semi-nested RT-PCR was performed using primers based on 312–314-base pair (bp) sequences in the ORF1–2 junction region [6]. For genotypic confirmation, VP1 complete nucleotide sequences were analyzed along with four representative outbreak cases that occurred in relatively large cities in Korea. In brief, cDNA was synthesized using SuperScript™ III Reverse Transcriptase (Invitrogen, Carlsbad, CA, USA) according to the manufacturer’s instructions and used as template DNA. Forward and reverse primers (GII17F, ACCATGAAGACCCCAGTGAG; GII17R, CCTGCCAATCCTGCAATGAA) were designed based on the complete genome of GII.17-Kawasaki323 (GenBank accession no. AB983218). The PCR conditions were: initial denaturation for 5 minutes at 95°C, annealing for 1 minutes at 50°C, extension for 3 minutes at 72°C, and final extension for 7 minutes at 72°C.
MATERIALS AND METHODS
- VP1 complete nucleotide sequence homology between the four representative specimens (Seoul, Incheon, Gyeonggi, and Daegu) was determined as follows. None of these specimens shared epidemiological relationships, but were representative patient samples from an outbreak in each region. Nucleotide sequence homology between representative specimens ranged from 98.3% to 99.9%, and detailed information is shown in Table 1.
- The sequenced partial capsid gene was identified as GII.17 using K-EnterNet, the NoroNet typing tool, and a nucleotide BLAST. The pairwise distances of amino acid sequences also aligned with those of other GII.17 strains on NoroNet (accession no. DQ38972, AY502009) and the Kawasaki323 strain (accession no. AB983218) in an analysis using the CLUSTAL W program in the MegAlign package (Windows version 3.12e). However, prevalent GII.17 strains in Korea differed from those strains by 4.4% (AB AB983218), 13.0% (DQ38972), and 10.9% (AY502009). The relationships of nucleotide sequences are shown in a den-drogram and in phylogenetic trees generated using the neighbor-joining method and based on the NoroNet GII reference set and relatively recently identified GII.17 variants (Kawasaki323, Kawasaki308, Nagano7-1, and Saitama5203; Figure 1).
- Norovirus GII.4 was the prominent genotype in all cases. In the 2012–2013 season, GII.4 was the prominent genotype as a result of emergence of the GII.4 Sydney variant, whereas GII.17 was only a minor genotype in outbreak cases. In the 2013–2014 season, GII.4 remained overwhelmingly prominent among sporadic cases, whereas no major causative strain was identified among outbreak cases. However, the detection of GII.17 increased sharply in Korea beginning in December 2014, along with a drastic decrease in the prevalence of GII.4. More recently, GII.17 has been identified as a major contributing pathogen, and no additional GII.4 strains were identified in a February outbreak or sporadic cases from May (Figure 2).
RESULTS
- GII.4 was the most prevalent norovirus strain in outbreaks during the 2012–2013 seasons, but became a minor strain during the 2014–2015 season. During the earlier season, the GII.4 Sydney variant was emergent, and this information and previous cases have led us to expect that a new GII.4 variant, rather than GII.17, will be the next prominent strain. Interestingly, GII.17 became a major prominent strain in both outbreaks and sporadic cases during the 2014–2015 season. According to a previous report, GII.17 accounted for 76% of the norovirus contamination of surface water in Kenya [7]. In Korea, there was no investigation reporting groundwater contamination with norovirus GII.17 during 2002–2003 [8], but there have been reports detecting GII.17 in commercial food processing facilities (2007) and metropolitan Seoul (2008) [9,10]. Furthermore, this strain has been detected continuously in food-catering facilities and was identified as a major genotype, in addition to GII.4, in the Han River and Wang-suk River (2010) [11]. GII.17 was also detected in several outbreaks from 2012 to 2015, but was not identified in sporadic cases. A GII.17 variant strain was recently reported to be involved in outbreaks in Japan and China [12–14]. GII.17 is therefore a major contributor in norovirus-related outbreaks and sporadic cases, although the underlying reasons remain unknown [4]. It is impossible to know the persistence of this phenomenon. However, GII.17 was predominant during 2014–2015 season than GII.4 [14]. Changes in the patterns of molecular epidemiology are questionable.
- In Korea, norovirus genotyping is routinely performed using 312-bp sequences from partial capsid genes. Here, partial sequence genotypes were not determined via comparisons with reference sequences using the MegAlign program and NoroNet typing tool. A BLAST search identified several similar genes that had largely been registered since 2012: Kibera209 (KF916585, 2012), GyungGi88 (KF773972, 2012), and Kawasaki323 (AB983218, 2014). The Kawasaki323 strain was the only registered complete genome and exhibited 96% amino acid similarity with the emerging GII.17 strain in Korea. However, the Kawa-saki323 strain and emerging GII.17 strain in Korea differed from the NoroNet GII.17 reference genes by approximately 10.8% to 13.4%. This difference was near the cut-off value for distinguishing between genotypes [1] and might represent a limitation of norovirus genotype determination from partial VP1 sequences.
- Although we expect that a new GII.4 variant will emerge in Korea, the GII.17 strain is clearly major contributing genotype. Questions remain regarding the rapid genotypic shift from GII.4 to GII.17 in Korea. The following may provide an important clue. First, no dominant genotype was identified among outbreak cases during the 2013–2014 season. Although GII.4 was always the dominant genotype worldwide, it did not induce outbreaks during the 2013–2014 season. Second, although a significant proportion of sporadic cases were attributed to GII.4, the prevalence of GII.17 increased rapidly among both outbreak and sporadic cases since December 2014. Continuous monitoring and further investigations of environmental contamination are needed to answer the above question.
DISCUSSION
-
Acknowledgements
- This study was supported by the National Norovirus Surveillance System (K-CaliciNet) and the acute diarrheal laboratory surveillance system (K-EnterNet) (4851-304-201).
ACKNOWLEDGMENTS
-
CONFLICTS OF INTEREST
No potential conflict of interest relevant to this article was reported.
Article information
- 1. Kroneman A, Vega E, Vennema H, et al. Proposal for a unified norovirus nomenclature and genotyping. Arch Virol 2013;158:2059−68. https://doi.org/10.1007/s00705-013-1708-5. PMID: 10.1007/s00705-013-1708-5. PMID: 23615870.ArticlePubMedPMC
- 2. Park K, Yeo S, Jeong H, et al. Updates on the genetic variations of norovirus in sporadic gastroenteritis in Chungnam Korea, 2009–2010. Virol J 2012;9:29https://doi.org/10.1186/1743-422X-9-29. PMID: 10.1186/1743-422X-9-29. PMID: 22273062.ArticlePubMedPMC
- 3. Cho HG, Lee SG, Kim JE, et al. Molecular epidemiology of norovirus GII.4 variants in children under 5 years with sporadic acute gastro-enteritis in South Korea during 2006–2013. J Clin Virol 2014;61:340−4. https://doi.org/10.1016/j.jcv.2014.08.018. PMID: 10.1016/j.jcv.2014.08.018. PMID: 25223918.ArticlePubMed
- 4. de Graaf M, van Beek J, Vennema H, et al. Emergence of a novel GII.17 norovirus – End of the GII.4 era? Euro Surveill 2015;20:https://doi.org/10.2807/1560-7917.ES2015.20.26.21178. PMID: 10.2807/1560-7917.ES2015.20.26.21178.Article
- 5. Korea Centers for Disease Control and Prevention. Guideline for water and foodborne diseases prevention and control. Chungju: Korea Centers For Disease Control and Prevention; 2016.
- 6. Kim SH, Cheon DS, Kim JH, et al. Outbreaks of gastroenteritis that occurred during school excursions in Korea were associated with several waterborne strains of norovirus. J Clin Microbiol 2005;43:4836−9. https://doi.org/10.1128/JCM.43.9.4836-4839.2005. PMID: 10.1128/JCM.43.9.4836-4839.2005. PMID: 16145153.ArticlePubMedPMC
- 7. Kiulia NM, Mans J, Mwenda JM, et al. Norovirus GII.17 predominates in selected surface water sources in Kenya. Food Environ Virol 2014;6:221−31. https://doi.org/10.1007/s12560-014-9160-6. PMID: 10.1007/s12560-014-9160-6. PMID: 25059212.ArticlePubMed
- 8. Lee C, Kim SJ. The genetic diversity of human noroviruses detected in river water in Korea. Water Res 2008;42:4477−84. https://doi.org/10.1016/j.watres.2008.08.003. PMID: 10.1016/j.watres.2008.08.003. PMID: 18778846.ArticlePubMed
- 9. Jung JH, Yoo CH, Koo ES, et al. Occurrence of norovirus and other enteric viruses in untreated groundwaters of Korea. J Water Health 2011;9:544−55. https://doi.org/10.2166/wh.2011.142. PMID: 10.2166/wh.2011.142. PMID: 21976201.ArticlePubMed
- 10. Lee H, Kim M, Lee JE, et al. Investigation of norovirus occurrence in groundwater in metropolitan Seoul, Korea. Sci Total Environ 2011;409:2078−84. https://doi.org/10.1016/j.scitotenv.2011.01.059. PMID: 10.1016/j.scitotenv.2011.01.059. PMID: 21440930.ArticlePubMed
- 11. Lee BR, Lee SG, Park JH, et al. Norovirus contamination levels in ground water treatment systems used for food-catering facilities in South Korea. Viruses 2013;5:1646−54. https://doi.org/10.3390/v5071646. PMID: 10.3390/v5071646. PMID: 23820792.ArticlePubMedPMC
- 12. Lu J, Sun L, Fang L, et al. Gastroenteritis outbreaks caused by norovirus GII.17, Guangdong province, China, 2014–2015. Emerg Infect Dis 2015;21:1240−2. https://doi.org/10.3201/eid2107.150226. PMID: 10.3201/eid2107.150226. PMID: 26080037.ArticlePubMedPMC
- 13. Fu J, Ai M, Jin J. Emergence of a new GII.17 norovirus variant in patients with acute gastroenteritis in Jiangsu, China, september 2014 to march 2015. Euro Surveill 2015;20:https://doi.org/10.2807/1560-7917.ES2015.20.24.21157. PMID: 10.2807/1560-7917.ES2015.20.24.21157. PMID: 26111236.Article
- 14. Matsushima Y, Ishikawa M, Shimizu T, et al. Genetic analyses of GII.17 norovirus strains in diarrheal disease outbreaks from December 2014 to March 2015 in Japan reveal a novel polymerase sequence and amino acid substitutions in the capsid region. Euro Surveill 2015;20:https://doi.org/10.2807/1560-7917.ES2015.20.26.21173. PMID: 10.2807/1560-7917.ES2015.20.26.21173. PMID: 26159307.Article
REFERENCES
Figure 1
Neighbor-joining phylogenetic tree based on complete VP1 nucleotide sequences (located in ORF2) detected in outbreak cases. Reference sequences were collected from NoroNet and GenBank. Numbers in branches indicate the bootstrap values and outbreak strains classified as genogroup (G) II.17.
SE, Seoul; GG, Gyeonggi; DG, Daegu; IC, Incheon.
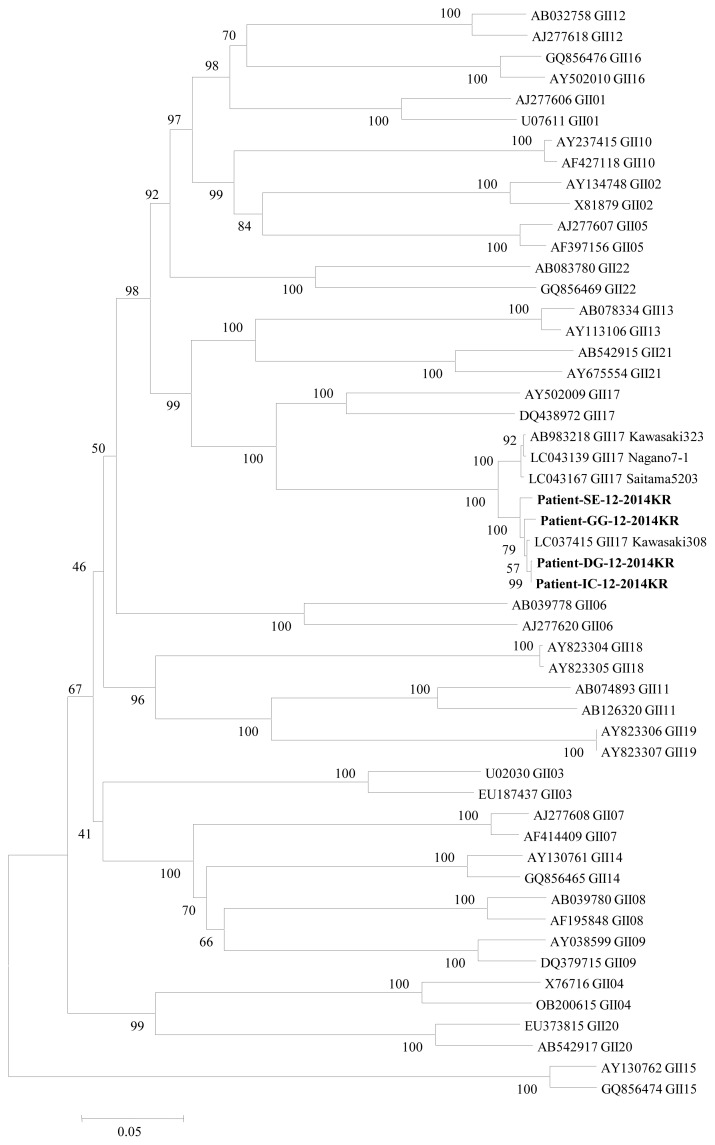


Figure 2Prevalence of norovirus genotypes in Korea from 2012 to 2014. A total of 4,964 partial VP1 sequences were used to analyze genotypes and separate them into three groups: genogroup (G) II.4, GII.17, and other norovirus types. (A) Prevalence of norovirus genotypes among outbreaks. The prevalence of GII.17 increased explosively since December 2014. (B) Prevalence of norovirus genotypes among sporadic cases. The GII.17 strain, which had not previously appeared, increased in prevalence since December 2014.
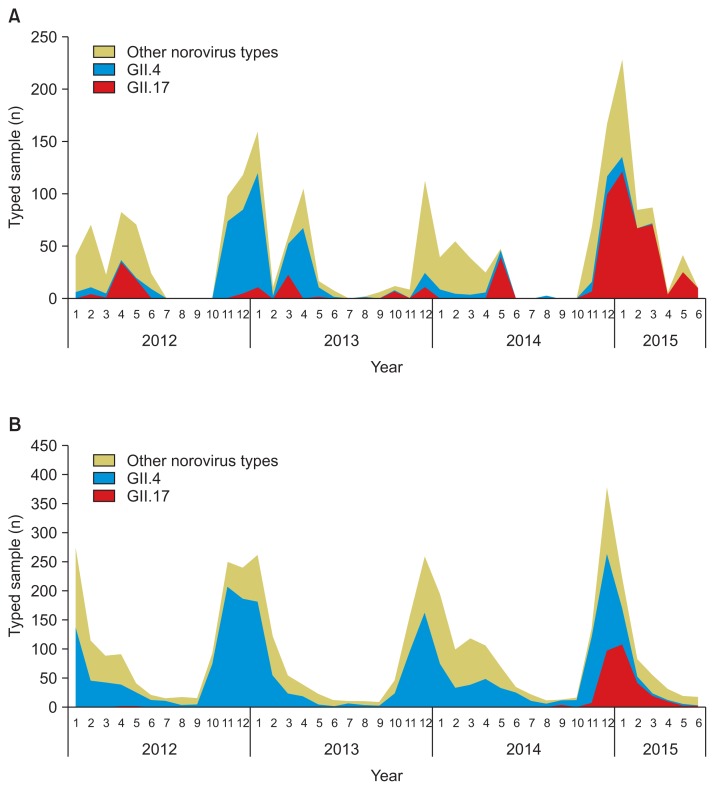

Table 1Nucleotide sequence homology of GII.17 complete VP1 sequences of representative specimens from Seoul (SE), Incheon (IC), Gyeonggi (GG), and Daegu (DG) in Korea (KR) (%)
Figure & Data
References
Citations
Citations to this article as recorded by 

- Monitoring of foodborne viruses in pre- and post-washed root vegetables in the Republic of Korea
Sunho Park, Md Iqbal Hossain, Soontag Jung, Zhaoqi Wang, Daseul Yeo, Seoyoung Woo, Yeeun Seo, Myeong-In Jeong, Changsun Choi
Food Control.2023; 154: 109982. CrossRef - Epidemiology of Norovirus Outbreaks Reported to the Public Health Emergency Event Surveillance System, China, 2014–2017
Yiyao Lian, Shuyu Wu, Li Luo, Bin Lv, Qiaohong Liao, Zhongjie Li, Jeanette J. Rainey, Aron J. Hall, Lu Ran
Viruses.2019; 11(4): 342. CrossRef - Nearly Complete Genome Sequence of a Human Norovirus GII.P17-GII.17 Strain Isolated from Brazil in 2015
Cristina Santiso-Bellón, Azahara Fuentes-Trillo, Juliana da Silva Ribeiro de Andrade, Carolina Monzó, Susana Vila-Vicent, Roberto Gozalbo Rovira, Javier Buesa, Felipe J. Chaves, Marize Pereira Miagostovich, Jesús Rodríguez-Díaz, Jelle Matthijnssens
Microbiology Resource Announcements.2019;[Epub] CrossRef - Development of SNP Markers for Norovirus Related FUT2 Gene in Oyster
守泉 闫
Hans Journal of Biomedicine.2019; 09(03): 135. CrossRef - Research on the contamination levels of norovirus in food facilities using groundwater in South Korea, 2015–2016
Jeong Su Lee, In Sun Joo, Si Yeon Ju, Min Hee Jeong, Yun-Hee Song, Hyo Sun Kwak
International Journal of Food Microbiology.2018; 280: 35. CrossRef - Norovirus GII.17 Associated with a Foodborne Acute Gastroenteritis Outbreak in Brazil, 2016
Juliana da Silva Ribeiro de Andrade, Tulio Machado Fumian, José Paulo Gagliardi Leite, Matheus Ribeiro de Assis, Alexandre Madi Fialho, Sergio Mouta, Cristiane Mendes Pereira Santiago, Marize Pereira Miagostovich
Food and Environmental Virology.2018; 10(2): 212. CrossRef - Phylogenetic characterization of norovirus strains detected from sporadic gastroenteritis in Seoul during 2014–2016
Young Eun Kim, Miok Song, Jaein Lee, Hyun Jung Seung, Eun-Young Kwon, Jinkyung Yu, Youngok Hwang, Taeho Yoon, Tae Jun Park, In Kyoung Lim
Gut Pathogens.2018;[Epub] CrossRef - The prevalence of non-GII.4 norovirus genotypes in acute gastroenteritis outbreaks in Jinan, China
Lanzheng Liu, Hengyun Guan, Ying Zhang, Chunrong Wang, Guoliang Yang, Shiman Ruan, Huailong Zhao, Xiuyun Han, Adriana Calderaro
PLOS ONE.2018; 13(12): e0209245. CrossRef


 PubReader
PubReader Cite
Cite
