Articles
- Page Path
- HOME > Osong Public Health Res Perspect > Volume 3(1); 2012 > Article
-
Articles
Viral Interferon Regulatory Factor 1 of Kaposi’s Sarcoma-Associated Herpesvirus Interacts with a Translocation Liposarcoma Protein-Associated Serine-Arginine Protein - Sunmi Kim, Jae Eun Jong, Taegun Seo
-
Osong Public Health and Research Perspectives 2011;3(1):8-13.
DOI: https://doi.org/10.1016/j.phrp.2012.01.001
Published online: December 31, 2011
Department of Life Science, Dongguk University-Seoul, Seoul, Korea.
- Corresponding author. E-mail: tseo@dongguk.edu
• Received: December 8, 2011 • Revised: January 5, 2012 • Accepted: January 15, 2012
Copyright ©2012, Korea Centers for Disease Control and Prevention
This is an Open Access article distributed under the terms of the Creative Commons Attribution Non-Commercial License () which permits unrestricted non-commercial use, distribution, and reproduction in any medium, provided the original work is properly cited.
- 2,442 Views
- 12 Download
Abstract
-
Objectives
- To confirm that Kaposi’s sarcoma-associated herpes virus openreading frame K9, viral interferon regulatory factor 1 (vIRF1), interacts with splicing factor, translocation liposarcoma protein-associated serine-arginine protein (TASR), in vivo and to establish whether interactions between vIRF1 and TASRs influence alternative splicing.
-
Methods
- Association between vIRF1 and TASRs was confirmed with the glutathione S-transferase pull-down assay and coimmunoprecipitation. Further colocalization was shown by immunofluorescence. The in vivo splicing assay was performed to confirm the alterations in the splicing pattern.
-
Results
- vIRF1 interacts with both TASR1 and 2 in vivo. vIRF1 has been shown to colocalize with TASR proteins in 293 T cells. However, an in vivo splicing revealed no alterations in the splicing pattern via interaction.
-
Conclusions
- The study data suggest that vIRF1 interacts with the TASR protein. However, vIRF1 interactions do not affect TASR-mediated alternative splicing.
- Kaposi’s sarcoma-associated herpesvirus (KSHV), also known as human herpes virus 8, is a gammaherpesvirus. Epstein-Barr virus (EBV) and saimiri herpesvirus belong to this virus group and display similar genetic properties. KSHV triggers B cell lymphoma, primary effusion lymphoma and multicentric Castleman’s disease [1-3]. When the host cell is infected with a virus, interferons (IFN) are activated as part of the immune response, followed by triggering of various biological activities, including suppression of tumor, apoptosis regulated by the cell cycle, and immune activation. IFN activation is regulated by IFN regulatory factors (IRF) that recognize the IFN-stimulated response element (ISRE) [4-8]. KSHV has four open reading frames (ORF) encoding viral IRFs (vIRF) that are homologous to the cellular IRF family [3,4,9-11].
- A number of studies show that vIRF1 acts as a transcriptional activator, while others have reported that vIRF4 interacts with and inhibits transactivation of various cellular factors [7]. For example, the tumor suppressor p53 interacts with vIRF1, leading to repression of the transcriptional activation of p53, and consequently, inhibition of p53-dependent apoptosis [12,13]. vIRF1 additionally interacts with GRIM19, a cell death regulator, and inhibits GRIM19-mediated apoptosis in the presence of IFN/retinoic acid [14]. Moreover, malignant tumors have been generated in nude mice injected with NIH3T3 cells stably expressing vIRF1 [4]. These studies indicate that vIRF1 influences KSHV tumorigenicity as a viral oncogene. vIRF1 also interacts with the transcriptional coactivator, p300/ CREB, and inhibits transcription of the IFNA gene by suppressing the transactivation of cellular IRFs. Earlier studies have additionally reported that vIRF1 suppresses cellular cytokine expression by inhibiting the histone acetyltransferase activity of p300 [9,15,16]. Thus, vIRF1 clearly alters the expression patterns of cellular factors.
- The serine-arginine (SR) protein family is important for pre-mRNA splicing. These proteins contain two major domains. The N-terminus of SR proteins consists of ribonucleoprotein-type RNA binding domains that interact with pre-mRNA, and the C-terminus includes an arginine-serine (RS) domain required for protein-protein interactions [17,18].
- The SR splicing factors, translocation liposarcoma protein (TLS)-associated serine-arginine proteins (TASRs), have been classified into two isoforms, TASR 1 and 2, composed of 183 and 262 aa, respectively. The proteins share a common N-terminal domain, but differ in the C-terminal domain. An earlier in vivo E1A splicing assay revealed differential splicing patterns. TASR 1 induces splicing to the 11S and 10S isoforms, and TASR2 to the 9S isoform [19-21]. Additionally, TASR 1 contributes to alternative splicing of type II and XI collagen genes in chondrogenic cells, but TASR2 does not [22]. TASRs interact with TLS or Ewing’s sarcoma (EWS) proteins through the C-terminus. Normally, TLS and EWS link the gene transcription of RNA polymerase II to RNA splicing of TASRs. Upon replacement of the C-terminus with ETS-related gene and Friend leukemia integration-1, alternative splicing through TASRs was disrupted, and human myeloid leukemia and sarcoma were triggered, respectively [20,21,23-25].
- Previously, we identified an association between TASRs and vIRF1 using the yeast two-hybrid assay [14]. Interactions with vIRF1 potentially influence the functions of cellular proteins, and we therefore assumed that vIRF1 affects the splicing function of TASRs via binding. However, the results indicate that vIRF1 does not affect the splicing function of TASRs.
1. Introduction
- 2.1. Plasmids
- To construct Flag-tagged TASR1 and TASR2, TASR cDNA was amplified from pcDNA3-TASR1 and pcDNA3-TASR2, respectively. The forward TASR primer, 5' CCC GAA TTC ATG TCC CGC TAC CTG CGT 3', reverse TASR1 primer, 5' GGG CTC GAG TCA GTGGCCACTGGACTT 3', and reverseTASR2 primer, 5' GGG CTC GAG TCA GAT CTT TCT TGA AGT GTA 3', were employed for amplification. Sequences were subcloned into the EcoRI/XhoI site of the Flagtagged vector, pME18S.
- 2.2. Cell culture and transfection
- 293 T and COS-1 cells were grown in Dulbecco’s modified Eagle’s medium containing 10% fetal bovine serum and 1% antibiotics. For the binding assay, 293 T cells were cotransfected with pEBG or pEBG-vIRF1 and pME18S-TASR1 or pME18S-TASR2 using the calcium phosphate method. For the immunofluorescence assay, 293 T cells were cotransfected with pEGFP-C1- vIRF1 and pME18S-TASR using PolyExpress reagents (Excellgen, Rockville, MD). COS-1 cells were transfected with pCS3-MT-E1A, pME18S-TASRs, and pcDNA3-vIRF1, using the same reagents.
- 2.3. Glutathione S-transferase pull-down assay and western blotting
- At 48 h posttransfection, 293 T cells were lysed in EBC buffer (50 mM Tris-HCl [pH7.5], 120 mM NaCl, 50 mM NaF, 0.5% Nonidet P-40, 1 mM phenylmethylsulfonyl fluoride). Cell lysates were incubated with glutathione sepharose 4B beads (Amersham Pharmacia Biotech, Cardiff, UK) for 2 h at 4℃. Glutathione Stransferase (GST) fusion protein and GST bead complexes were washed with EBC buffer. Samples were mixed with loading buffer, heated at 100℃, separated with SDS-PAGE on a 10% gel, and transferred to PVDF membrane. Proteins were immunoblotted with a monoclonal mouse anti-Flag antibody (Sigma-Aldrich, Poole, United Kingdom), and bands detected with the enhanced chemiluminescence analysis system (Thermo Scientific, Woburn, MA, USA).
- 2.4. Coimmunoprecipitation
- Transfected 293 T cells were lysed in EBC buffer at 48 h posttransfection. Lysates were centrifuged at 2440g for 20 min. Supernatant fractions were initially incubated with monoclonal mouse anti-Flag antibody for 1 h at 4℃ with rocking, and subsequently with protein G Sepharose 4 Fast Flow (GE Healthcare, Waukesha, WI, USA) beads for 1 h at 4℃. Flag fusion protein and bead complexes were washed with EBC buffer 4 times. Samples were mixed with loading buffer and heated at 100℃ for SDS-PAGE. Proteins were separated using SDS-PAGE, and immunoblotted with an anti-GST antibody.
- 2.5. Immunofluorescence
- Transfected 293 T cells were fixed with 3.7% formaldehyde for 15 min, and permeabilized with phosphatebuffered saline (PBS) containing 0.2% Triton X-100 (PBST) for 10 min. Cells were blocked with PBST containing 1% bovine serum albumin for 30 min, incubated with anti-Flag antibodies for 1 h, and washed 3 times with PBST. Next, cells were incubated with rhodamineconjugated anti-mouse IgG antibodies for 1 h, washed 3 times with PBST, and analyzed using confocal microscopy (FV-1000; Olympus, Tokyo, Japan).
- 2.6. E1A pre-mRNA splicing assay
- COS-1 cells were cotransfected with 0.25 mg of pCS3-MT-E1A, 1.5 mg of pcDNA3-vIRF1 and 0.25 mg of pME18S-TASR1 or 2 using PolyExpress reagents. The total amount of DNA was adjusted with corresponding empty vectors. After 24 h, total RNA was extracted with TRIzol reagent and reverse-transcribed
- with oligo(dT) primers. Various E1a splice forms were amplified with these primers. The forward primer used was 5'-GAG CTT GGG CGA CCT CA-3' (RR67) and reverse primer was 5'-TCT AGA CAC AGG TGA TGT CG-3' (E1A2). PCR products were separated on a 1.5% agarose gel containing ethidium bromide and analyzed with gel documentation.
2. Materials and Methods
Figure 1.
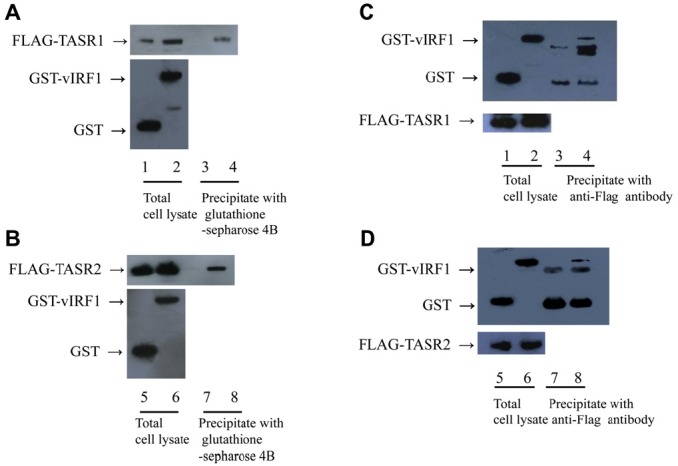
Interactions of vIRF1 with TASR1 or 2 in vivo. (A) 293 T cells were cotransfected with GST or GST-vIRF1 and Flag- TASR1 expression plasmids. Cells were lysed with EBC buffer, and cell extracts incubated with glutathione sepharose 4B beads. GST fusion and Flag fusion proteins were detected using western blotting with anti-GST (top panel) and anti-Flag antibodies (bottom panel), respectively. Lanes 1, 3, GST and Flag-TASR1; 3, 4, GST-vIRF1 and Flag-TASR1 (B) 293 T cells were cotransfected with GST or GST-vIRF1 and Flag-TASR2 expression plasmids and subjected to similar experiments as those specified in Panel A. Lanes 5, 7, GST and Flag-TASR2; Lanes 6, 8, GST-vIRF1 and Flag-TASR2. (C) Reciprocal assay for Panel A. 293 T cells were cotransfected with GST or GST-vIRF1 and Flag-TASR1 expression plasmids. Cell extracts were incubated with anti-Flag antibody and precipitated with protein G-Sepharose beads for 1 h. GST fusion and Flag fusion proteins were detected by western blotting with anti-GST and anti-Flag antibodies respectively. Lanes 1, 3, GST and Flag-TASR1; Lanes 2, 4, GST-vIRF1 and Flag-TASR1 (D) Reciprocal assay for Panel B. 293 T cells were cotransfected with GST or GST-vIRF1 expression plasmid and Flag-TASR2, and subjected to similar experiments as those specified in panel C. Lanes 5, 7, GST and Flag-TASR1; Lanes 6, 8, GST-vIRF1 and Flag-TASR1.

- 3.1. vIRF1 interacts with TASR1 and TASR2 in vivo
- Having previously confirmed the association of vIRF1 and TASR1 using the yeast two-hybrid assay [14], we aimed to further ascertain whether TASR2 also interacts with vIRF1, since both isoforms contain identical Nterminal domains. To identify the in vivo interactions, 293 T cells were cotransfected with GST or GST-vIRF1, and Flag-TASR1 or Flag-TASR2 expression plasmids. At 48 h posttransfection, proteins were extracted from transfected cells, and GST and GST-vIRF1 precipitated with glutathione sepharose 4B beads. Proteins were separated via SDS-PAGEand thewestern blot assay performed using an anti-Flag antibody. Both Flag-TASR1 and Flag-TASR2 coprecipitated with GST-vIRF1 (Figure 1A, Lane 4, and Figure 1B, Lane 8), but not GST alone (Figure 1A, Lane 3, Figure 1B, Lane 7). To verify the correct expression of GST, GST-vIRF1, Flag-TASR1, and Flag-TASR2 proteins, whole-cell lysates were immunoblotted using anti-GST and anti-Flag antibodies (Figure 1A Lane 1, 2 and Figure 1B Lane 5, 6). In a reciprocal assay, cell lysates were incubated with anti-Flag antibody for 1 h, followed by protein G-Sepharose beads for 1 h. Precipitated proteins were separated by SDS-PAGE and immunoblotted using an anti-GSTantibody.GST-vIRF1was coprecipitated with Flag-TASR1 and Flag-TASR2 (Figure 1C, Lane 4, and Figure 1D, Lane 8). To establish whether proteins were expressed correctly, western blot assay for total lysates was performed (Figure 1C, Lanes 1, 2, and Figure 2C, Lanes 5, 6). Our results indicate that vIRF1 interacts with both TASR1 and TASR2 in mammalian cells.
- 3.2. vIRF1 colocalizes with TASR1 and TASR2
- To confirm colocalization of vIRF1 with TASRs, we performed immunofluorescence assay. Green fluorescent protein (GFP)-vIRF1 proteins were predominantly expressed in the nucleus, while GFP was expressed in
- nucleus and cytoplasm. Flag-TASR1 and 2 were mostly localized in the nucleoplasm. 293 T cells were further cotransfected with GFP-vIRF1 and Flag-TASR1 or 2 expression plasmids, and the expression patterns analyzed using confocal microscopy. Notably, vIRF1 colocalized with TASR1 and 2 in the nucleus, particularly within the nucleoplasm region (Figure 2).
- 3.3. vIRF1 has no effect on E1A mRNA splicing mediated by TASR1 and 2
- TASR1 and 2 are known splicing factors. In view of the finding that TASR1 and 2 interact with vIRF1, we subsequently examined whether vIRF1 affects the splicing function of TASRs. The well-characterized E1A reporter gene is commonly employed for analysis of splicing. E1A pre-mRNA exists as five spliced isoforms (Figure 3A). Cos-1 cells were transfected with various combinations of E1A reporter plasmids, TASRs, and vIRF1 (Figure 3B, top). TotalRNAwas extracted, andRT-PCRperformed for amplification of the E1A pre-mRNA isoforms. COS-1 cells mainly transfected with E1A reporter plasmids generated 13S, 12S, and 9S isoforms (Figure 3B Lane 1). Overexpression of TASR1 induced the generation of 11S, 10S and 9S, while TASR2 promoted an increase in the 9S isoform (Figure 3B, Lanes 3 and 5). Coexpression of vIRF1 did not affect TASR-mediated E1A mRNA splicing (Figure 3B, Lanes 4 and 6).
3. Results
Figure 2.

Colocalization of vIRF1 and TASR1 or 2. (A) GFP, GFP-vIRF1, Flag-TASR1 and Flag-TASR2 were transfected in 293 T cells, respectively. Cells were fixed at posttransfection 24 h. Flag-TASR1/2-expressing cells were incubated with anti-Flagantibody and detected with TRITC-conjugated anti-mouse-antibody. (B) Colocalization of vIRF1 and TASR1. Both vIRF1 and Flag-TASR1 were localized in the nucleus. (C) Colocalization of vIRF1 and TASR2 (x 600 magnification).

Figure 3.
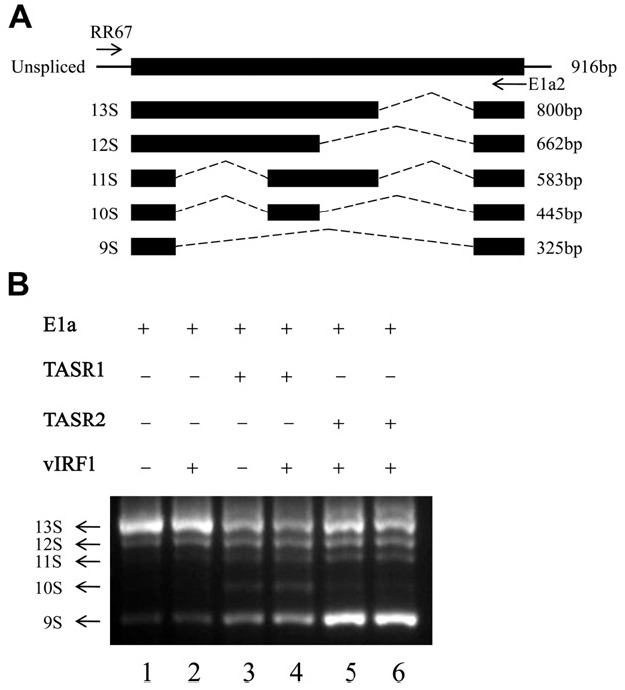
In vivo E1A splicing assay. (A) Diagram of E1A pre-mRNA splicing isoforms. RT-PCR primers are represented by arrows (RR67: forward primer, E1a2: reverse primer). (B) In vivo E1A splicing assay. COS-1 cells were cotransfected with pCS3-MT-E1A, pcDNA3-vIRF1 and pME18S-TASR1 or 2. After 24 h, total RNA was extracted and reverse-transcribed using oligo(dT) primers. E1A pre-mRNA splicing isoforms were amplified with a pair of specific primers, as described in Panel A.

- KSHV proteins are significantly homologous to host cellular proteins, and may be essential to avoid host immune surveillance activated by viral infections [6]. In particular, vIRF1 (ORF9), which is homologous to cellular IRF, inhibits transcriptional processes regulated by types I and II IFN and the tumor suppressor, IRF1 [4,26]. Moreover, vIRF1 interacts with p53 and GRIM19, leading to the prevention of p53-dependent and GRIM19- induced apoptosis, respectively. Based on these results, we assumed that vIRF1 affects the tumorigenicity of KSHV as a repressor of cell cycle regulation [12-14].
- We identified that TASR1 interacts with vIRF1 using the yeast two-hybrid assay (data not shown). Both TASR1 and TASR2 displaying sequence homology to TASR1 interacted with vIRF1 in vivo (19). TASR1 and 2 colocalized with vIRF1 in the nucleus. TASRs, initially identified from a yeast two-hybrid assay for TLS, are recruited to TLS or EWS. In human myeloid leukemia and malignant liposarcoma, the TLS C-terminus is replaced with ERG or CHOP fusion protein, respectively. Thus, SR proteins cannot be recruited to TLS, and SR-mediated E1A pre-mRNA splicing is effectively prevented [20,21]. We assume that if vIRF1 inhibits the splicing function of TASRs, interactions between these proteins should affect tumorigenicity, since both vIRF1 and TASR1 can induce tumors. Accordingly, we aimed to establish whether interactions between vIRF1 and TASRs influence alternative splicing. An in vivo splicing assay was performed to clarify this issue. In contrast to our expectation, vIRF1 did not appear to alter the TASRmediated splicing site selection via inhibition of TASR function. In case of effects on splicing, vIRF1 should be able to regulate the host system at the posttranscriptional level. However, vIRF1-TASR interactions did not influence RNA processing.
4. Discussion
-
Acknowledgements
- This research was supported by Basic Science Research Program through the National Research Foundation of Korea (NRF) funded by the Ministry of Education, Science and Technology (2009-0074240).
- 1. Soulier J Grollet L Oksenhendler E et al. Kaposi’s sarcomaassociated herpesvirus-like DNA sequences in multicentric Castleman’s disease. Blood 8;1995;86(4). 1276−80. PMID: 7632932.ArticlePubMed
- 2. Chang Y Cesarman E Pessin MS et al. Identification of herpesvirus-like DNA sequences in AIDS-associated Kaposi’s sarcoma. Science 12;1994;266(5192). 1865−9. PMID: 7997879.ArticlePubMed
- 3. Russo JJ Bohenzky RA Chien MC et al. Nucleotide sequence of the Kaposi sarcoma-associated herpesvirus (HHV8). Proc Natl Acad Sci U S A 1996;93(25). 14862−7. PMID: 8962146.ArticlePubMedPMC
- 4. Gao SJ Boshoff C Jayachandra S et al. KSHV ORF K9 (vIRF) is an oncogene which inhibits the interferon signaling pathway. Oncogene 10;1997;15(16). 1979−85. PMID: 9365244.ArticlePubMed
- 5. Li M Lee H Guo J et al. Kaposi’s sarcoma-associated herpesvirus viral interferon regulatory factor. J Virol 7;1998;72(7). 5433−40. PMID: 9620998.ArticlePubMedPMC
- 6. Ploegh HL . Viral strategies of immune evasion. Science 4;1998;280(5361). 248−53. PMID: 9535648.ArticlePubMed
- 7. Rezaee SA Cunningham C Davison AJ et al. Kaposi’s sarcomaassociated herpesvirus immune modulation: an overview. J Gen Virol 7;2006;87(Pt 7). 1781−804. PMID: 16760382.ArticlePubMed
- 8. Kirchhoff S Schaper F Hauser H . Interferon regulatory factor 1 (IRF-1) mediates cell growth inhibition by transactivation of downstream target genes. Nucleic Acids Res 6;1993;21(12). 2881−9. PMID: 8332497.ArticlePubMedPMC
- 9. Burysek L Yeow WS Lubyová B et al.. Functional analysis of human herpesvirus 8-encoded viral interferon regulatory factor 1 and its association with cellular interferon regulatory factors and p300. J Virol 9;1999;73(9). 7334−42. PMID: 10438822.ArticlePubMedPMC
- 10. Park J Lee MS Yoo SM et al.. Identification of the DNA sequence interacting with Kaposi’s sarcoma-associated herpesvirus viral interferon regulatory factor 1. J Virol 11;2007;81(22). 12680−4. PMID: 17855527.ArticlePubMedPMC
- 11. Burysek L Pitha PM. . Latently expressed human herpesvirus 8- encoded interferon regulatory factor 2 inhibits double-stranded RNA-activated protein kinase. J Virol 3;2001;75(5). 2345−52. PMID: 11160738.ArticlePubMedPMC
- 12. Nakamura H Li M Zarycki J et al.. Inhibition of p53 tumor suppressor by viral interferon regulatory factor. J Virol 8;2001;75(16). 7572−82. PMID: 11462029.ArticlePubMedPMC
- 13. Seo T Park J Lee D et al.. Viral interferon regulatory factor 1 of Kaposi’s sarcoma-associated herpesvirus binds to p53 and represses p53-dependent transcription and apoptosis. J Virol 7;2001;75(13). 6193−8. PMID: 11390621.ArticlePubMedPMC
- 14. Seo T Lee D Shim YS et al.. Viral interferon regulatory factor 1 of Kaposi’s sarcoma-associated herpesvirus interacts with a cell death regulator, GRIM19, and inhibits interferon/retinoic acidinduced cell death. J Virol 9;2002;76(17). 8797−807. PMID: 12163600.ArticlePubMedPMC
- 15. Li M Damania B Alvarez X et al.. Inhibition of p300 histone acetyltransferase by viral interferon regulatory factor. Mol Cell Biol. 11;2000;20(21). 8254−63. PMID: 11027294.ArticlePubMedPMC
- 16. Lin R Genin P Mamane Y et al.. HHV-8 encoded vIRF-1 represses the interferon antiviral response by blocking IRF-3 recruitment of the CBP/p300 coactivators. Oncogene 2;2001;20(7). 800−11. PMID: 11314014.ArticlePubMed
- 17. Tacke R Manley JL. . Determinants of SR protein specificity. Curr Opin Cell Biol 6;1999;11(3). 358−62. PMID: 10395560.ArticlePubMed
- 18. Van der Houven van Oordt W Newton K Screaton GR et al.. Role of SR protein modular domains in alternative splicing specificity in vivo. Nucleic Acids Res 12;2000;15(28). 4822−31. PMID: 11121472.Article
- 19. Clinton JM Chansky HA Odell DD et al.. Characterization and expression of the human gene encoding two translocation liposarcoma protein-associated serine-arginine (TASR) proteins. Gene 6 2 2002;284(1-2). 141−7. PMID: 11891055.ArticlePubMed
- 20. Yang L Embree LJ Hickstein DD. . TLS-ERG leukemia fusion protein inhibits RNA splicing mediated by serine-arginine proteins. Mol Cell Biol. 5;2000;20(10). 3345−54. PMID: 10779324.ArticlePubMedPMC
- 21. Yang L Embree LJ Tsai S et al.. Oncoprotein TLS interacts with serine-arginine proteins involved in RNA splicing. J Biol Chem 10;1998;273(43). 27761−4. PMID: 9774382.ArticlePubMed
- 22. Matsushita H Blackburn ML Klineberg E et al.. TASR-1 regulates alternative splicing of collagen genes in chondrogenic cells. Biochem Biophys Res Commun 5;2007;356(2). 411−7. PMID: 17367759.ArticlePubMedPMC
- 23. Yang L Chansky HA Hickstein DD. . EWS.Fli-1 fusion protein interacts with hyperphosphorylated RNA polymerase II and interferes with serine-arginine protein-mediated RNA splicing. J Biol Chem 12;2000;275(48). 37612−8. PMID: 10982800.ArticlePubMed
- 24. Ichikawa H Shimizu K Hayashi Y et al.. An RNA-binding protein gene, TLS/FUS, is fused to ERG in human myeloid leukemia with t(16;21) chromosomal translocation. Cancer Res 6;1994;54(11). 2865−8. PMID: 8187069.PubMed
- 25. Kuroda M Ishida T Horiuchi H et al.. Chimeric TLS/FUS-CHOP gene expression and the heterogeneity of its junction in human myxoid and round cell liposarcoma. Am J Pathol 11;1995;147(5). 1221−7. PMID: 7485386.PubMedPMC
- 26. Zimring JC Goodbourn S Offermann MK. . Human herpesvirus 8 encodes an interferon regulatory factor (IRF) homolog that represses IRF-1-mediated transcription. J Virol 1;1998;72(1). 701−7. PMID: 9420276.ArticlePubMedPMC
Figure & Data
References
Citations
Citations to this article as recorded by 



 PubReader
PubReader Cite
Cite
