Articles
- Page Path
- HOME > Osong Public Health Res Perspect > Volume 11(5); 2020 > Article
-
Original Article
2019 Tabletop Exercise for Laboratory Diagnosis and Analyses of Unknown Disease Outbreaks by the Korea Centers for Disease Control and Prevention - Il-Hwan Kima, Jun Hyeong Janga, Su-Kyoung Joa, Jin Sun Noa, Seung-Hee Seoa, Jun-Young Kima, Sang-Oun Junga, Jeong-Min Kima, Sang-Eun Leea, Hye-Kyung Parkb, Eun-Jin Kimb, Jun Ho Jeona, Myung-Min Choia, Boyeong Ryub, Yoon Suk Jangb, Hwami Kimb, Jin Leeb, Seung-Hwan Shinb, Hee Kyoung Kimb, Eun-Kyoung Kimb, Ye Eun Parka, Cheon-Kwon Yooa, Sang-Won Leea, Myung-Guk Hana, Gi-Eun Rhiea, Byung Hak Kanga
-
Osong Public Health and Research Perspectives 2020;11(5):280-285.
DOI: https://doi.org/10.24171/j.phrp.2020.11.5.03
Published online: October 22, 2020
aCenter for Laboratory Control of Infectious Diseases, Korea Centers for Disease Control and Prevention, Cheongju, Korea
bCenter for Public Health Emergency Preparedness and Response, Korea Centers for Disease Control and Prevention, Cheongju, Korea
- *Corresponding author: Byung Hak Kang, Center for Laboratory Control of Infectious Diseases, Korea Centers for Disease Control and Prevention, Cheongju, Korea, E-mail: bhkang11@korea.kr
• Received: June 12, 2020 • Revised: July 30, 2020 • Accepted: August 20, 2020
Copyright ©2020, Korea Centers for Disease Control and Prevention
This is an open access article under the CC BY-NC-ND license (http://creativecommons.org/licenses/by-nc-nd/4.0/).
- 5,867 Views
- 106 Download
Abstract
-
Objectives
- The Korea Centers for Disease Control and Prevention has published “A Guideline for Unknown Disease Outbreaks (UDO).” The aim of this report was to introduce tabletop exercises (TTX) to prepare for UDO in the future.
-
Methods
- The UDO Laboratory Analyses Task Force in Korea Centers for Disease Control and Prevention in April 2018, assigned unknown diseases into 5 syndromes, designed an algorithm for diagnosis, and made a panel list for diagnosis by exclusion. Using the guidelines and laboratory analyses for UDO, TTX were introduced.
-
Results
- Since September 9th, 2018, the UDO Laboratory Analyses Task Force has been preparing TTX based on a scenario of an outbreak caused by a novel coronavirus. In December 2019, through TTX, individual missions, epidemiological investigations, sample treatments, diagnosis by exclusions, and next generation sequencing analysis were discussed, and a novel coronavirus was identified as the causal pathogen.
-
Conclusion
- Guideline and laboratory analyses for UDO successfully applied in TTX. Conclusions drawn from TTX could be applied effectively in the analyses for the initial response to COVID-19, an ongoing epidemic of 2019 – 2020. Therefore, TTX should continuously be conducted for the response and preparation against UDO.
- With the recent increase in the occurrence of emerging and variant infectious diseases, the risk of infectious disease outbreaks have increased. During the past 2 decades, around 20 different types of infectious diseases have emerged including severe acute respiratory syndrome (SARS; 2002) [1], influenza A caused by the H1N1 virus (2009) [2], Dabie bandavirus causing severe fever with thrombocytopenia syndrome (SFTS) (2009) [3], Middle East respiratory syndrome (MERS) (2012) [4], and febrile illness caused by the Alongshan virus (2017) [5]. Emerging infectious diseases will continue to occur in future and Coronavirus disease 2019 (COVID-19) is the latest infectious disease to cause a pandemic [6].
- To prepare against unknown disease outbreaks (UDO), the Korea Centers for Disease Control and Prevention (KCDC) published “A Guideline for Unknown Disease Outbreaks,” in 2019, which included laboratory diagnosis and analyses. The KCDC had been preparing for an outbreak of an infectious disease of unknown etiology since April 2018 when an algorithm for diagnosis and analysis was developed for each syndrome through the management of the Unknown Disease Outbreak Laboratory Analyses Task Force (UDOLATF). The KCDC had also been forming a panel list for diagnosis by exclusion, and a panel diagnostic kit for the exclusion test. Diseases were assigned a syndrome from a list of 5, including hemorrhagic fever, respiratory symptoms, neurological symptoms, a rash, and diarrhea. Since September 2019, the UDOLATF has been preparing for potential UDO using tabletop exercises (TTX), with a focus on laboratory diagnosis and analyses, based on “A Guideline for Unknown Disease Outbreaks.”
Introduction
- 1. Definition of UDO
- The term “unknown disease outbreaks” is described by the world health organization (WHO) as a cluster of outbreaks or diseases of unknown etiology [7]. The CDC, in the US have stated that an “unexplained respiratory disease outbreak” is a case where the causal pathogen of the respiratory disease cannot be determined [8]. The KCDC defines “unknown disease outbreaks” as “cases where an unidentified pathogen (cause) is spatially and temporally associated with a disease (including severe cases and death), leading to outbreaks” [9].
- 2. Response to UDO
- In 2019, the KCDC published a guideline in response to UDO. The guideline focused on the response to the occurrence or predicted occurrence of an infectious disease or a disease of unknown etiology. According to the guidelines, a health professional or the director of a medical institution may request an epidemiological investigation from the director of the KCDC in cases of UDO.
- The KCDC runs a surveillance system, which includes a patient based surveillance system such as the Emergency Department syndrome monitoring system, patient monitoring system (for severe acute respiratory infection and influenza-like illness), laboratory based monitoring system (influenza and respiratory viruses), and acute respiratory infectious network. A case based monitoring system is also in operation using the 1339 call-line or press release. When a request is made by local government for an epidemiological investigation for UDO, such as a system of surveillance to facilitate the determination of whether an investigation is appropriate for the case, there is discussion and case assessment, and decisions regarding the dispatch of an immediate response team, epidemiological investigation, and the required level of management.
- Once the decision to conduct an epidemiological investigation (through case assessment discussion) is made, a study is launched to determine an appropriate level of control for the site of the outbreak, with respect to patients, contact individuals, exposed individuals, medical staff, and site inspectors. Upon launching the study, access to the site of outbreak may be restricted for site preservation, and requests can be made for medical records or interviews of the medical staff.
- Laboratory diagnosis should primarily be conducted by the Research Institute of Public Health and Environment. If the local government is unable to perform the diagnosis, a request can be made to the KCDC after the relevant consultation. Regarding the laboratory diagnosis system for an unknown disease, the laboratory diagnosis algorithm for syndromes or the WHO guidelines on clinical sample collection should be referred to according to the given case [10]. If the cause of the unknown disease is pathogenic infection, the patient and follow-up management of the exposed individuals is carried out. If the unknown disease has been identified as non-infectious, the task is transferred to the relevant department or institution.
- 3. The UDO Laboratory Analyses Task Force
- The Unknown Disease Outbreak Laboratory Analyses Task Force (UDOLATF) has been in operation since April 2018. It consists of the relevant experts including microbiologists and virologists. Through more than 10 conferences and council meetings, the algorithms for diagnosis and analyses have been developed in preparation for potential UDO. A panel list for diagnosis by exclusion was formed, and a panel diagnostic kit for the exclusion tests are under development for the 5 syndromes including respiratory symptoms. In case of a respiratory syndrome, the panel list comprises of 17 species of bacteria and fungi, and 16 species of viruses. The panel diagnostic kit for the exclusion test is currently being developed and will enable the simultaneous detection of 33 species of microbes.
- 4. TTX exercises for laboratory analyses of UDO
- Based on the recent MERS and SARS outbreaks, the UDOLATF predicted UDO with respiratory symptoms caused by a novel coronavirus originating from abroad, and produced a scenario of the outbreak to conduct training for a rapid and effective response. The region of outbreak in the scenario was based on the regions in China where large-scale outbreaks have recently occurred because of a novel virus. Recently, various novel viruses have emerged in China including SARS-CoV causing SARS in 2002, Dabie bandavirus causing SFTS in 2011, and Alongshan virus causing tick-borne disease in 2016. In the past in Winnan, China, there was an outbreak of bubonic plague, which spread throughout Europe in the 14th century. The scenario for the training was produced based on a virtual, large-scale outbreak of an unexplained pneumonia in the Winnan region, which then spread to Korea. The scenario was completed following review meetings over a period of around 3 months and TTX, where specific execution plans for each stage, and response strategies for the scenario were discussed.
- 5. TTX scenario for unexplained respiratory disease outbreak
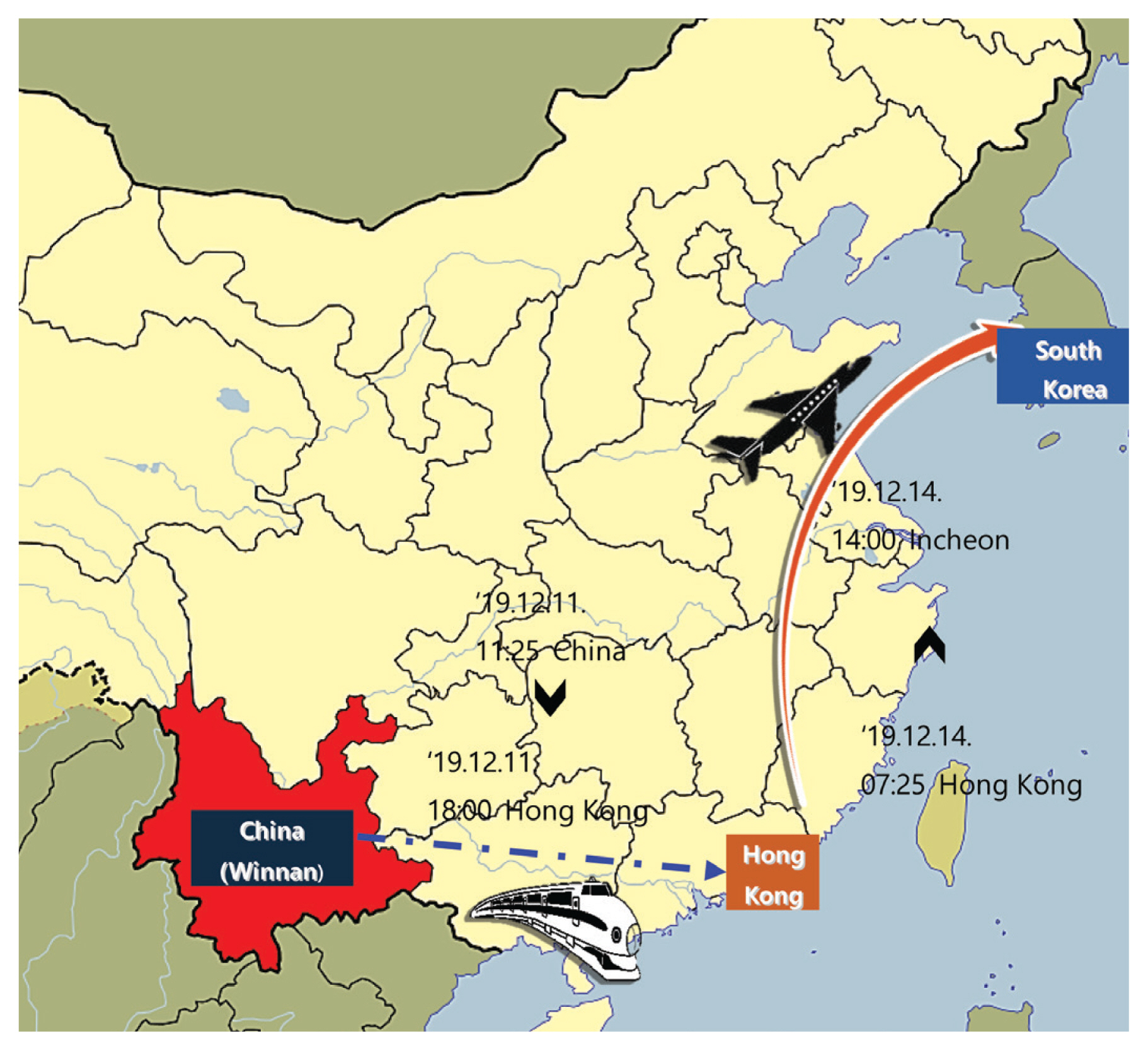
- A family of 4 Korean nationals [father #01 (41 years, male), mother #02 (40 years, female), and son #03 (16 years, male) and his grandfather #04 (66 years, male)] had been on a group tour to Winnan and Hong Kong in China. During the tour, the 4 people were together as a family, and they went mountain hiking in Winnan, then visited a cave, and a local farmer’s market (December 6th–11th, 2019), followed by a 3-day tour of Hong Kong (December 11th–13th, 2019), before returning to Korea via the Incheon Airport on December 14th, 2019 (Figure 1). After their return, symptoms began, and Patients #01 and #04 were admitted to a university hospital, where they passed away. The nurses at this hospital, where Patients #01, #02, and #04 were admitted, showed similar symptoms. There were 2 of the 6 individuals returning from the same package tour, had a fever (≥ 38°C) which was detected by thermal screening upon arriving at Incheon Airport.
- The clinical symptoms (Table 1) and the travel route of the family are as follows. During the evening before arrival at Incheon Airport, Patient #04 showed symptoms including a dry cough and runny nose. Upon arrival at the Incheon airport, Patients #01 and #02 began to show symptoms of a dry cough, and because no family member had a fever, they passed through the thermal screening at Incheon Airport undetected. The family returned to their home in Seoul in their own car.
- On Day 2 of arrival at dawn, Patient #04 had a fever and chills, whereas Patient #01 went to work in the morning in his own car. Symptoms including an itchy throat, slight cough, and mild fever began. Patient #01 joined a meeting with 10 colleagues at work in the afternoon and had dinner with 3 friends before returning home at 21:00.
- On Day 3 of arrival, Patient #04 suffered from aggravated symptoms of a fever (39.4°C), chills, and diarrhea at dawn. Patient #04 was taken to the Emergency Department of a university hospital by Patient #02 around 12:00, and was admitted. Whereas Patient #01 who went to work in his own car in the morning began to suffer from a severe cough, headache, fever (38.3°C), and myalgia, visited an internal medicine clinic close to his company, and was prescribed paracetamol for possible cold symptoms; he left work early. Patient #02, returned home from the Emergency Department after completing the admission process for Patient #04, where symptoms of fever and myalgia began later in the afternoon. Patient #01 took the prescribed paracetamol but still suffered from a fever of 38.5°C, a cough, chills, vomiting, and diarrhea; as Patient #02 showed similar aggravated symptoms of a fever, they visited the Emergency Department and were admitted for suspected acute pneumonia. Patient #03 showed no specific symptoms.
- On Day 4 of arrival, Patient #04 showed symptoms of a fever (40.1°C), ophthalmecchymosis, chest pain, dyspnea, and had convulsions at 13:00, followed by the symptoms of epistaxis, hemoptysis, and deterioration of consciousness, which aggravated until his death. At 18:00, Patient #01 showed similar symptoms as Patient #04, which aggravated until his death. At 18:30, Patient #02 was moved to the ICU because of dyspnea. The hospital where Patients #01, #02, and #04 were admitted, ran the tests for suspected diseases including respiratory and waterborne infectious diseases, but the causal pathogen could not be identified; a report was submitted to the KCDC Public Health Emergency Preparedness and Response Center via the 1339 call-line, and simultaneously the public health centers were alerted to the UDO. The request for an epidemiological investigation was made simultaneously to the KCDC, and the city of Seoul. At this time, the media in China and Hong Kong had reported on mortality cases due to an unidentified epidemic.
- On Day 5 at 10:20, 3 work colleagues of Patient #01 and 2 of his friends visited the Emergency Department (different regions) and were admitted due to similar symptoms as Patient #01. A nurse at the hospital where Patients #01, #02, and #04 were admitted to, began to show similar symptoms.
Materials and Methods
5.1. An overview of the unexplained respiratory disease outbreak cases
5.2. Clinical symptoms and the main travel route by the family
- 1. TTX for UDO
- On December 17th, 2019, TTX (using the cases presented) were performed by 22 participants, including the head of the KCDC Center for Laboratory Control of Infectious Disease, respective heads of the relevant departments, the epidemic intelligence service (EIS) officers, and laboratory analysis experts. The occurrence of UDO in the TTX scenario was confirmed by the EIS officers in charge at the KCDC Public Health Emergency Preparedness and Response Center. Following the case assessment discussion for the UDO, as well as the request for support to the city/province, it was decided that an immediate response team should be rapidly dispatched. The epidemiological investigator in the immediate response team (dispatched by the city of Seoul) quarantined the appropriate sites, and conducted an epidemiological investigation through interviews with medical staff, by review of the medical records, closed-circuit television, credit card records, and global positioning system data to analyze the detailed travel route of the deceased, and the infected patients, and the range of individuals the infected patients came into contact with.
- 2. The laboratory analyses for UDO
- The Laboratory Diagnosis Management division at the Center for Laboratory Control of Infectious Disease notified the other divisions of the occurrence of an infectious disease, and took charge of the liaison with the Public Health Emergency Preparedness and Response Center, and the task of relaying internal and external reports. The UDOLATF convened an emergency meeting to share the current information on the cases, and reviewed the laboratory diagnoses, testing methods, and the division of roles. The discussions among clinicians, medical practitioners, laboratory diagnosis experts, and EIS officers, and arbitration with the UDOLATF led to the determination of the sample collections, and subsequent test processes were reported to the head of the center at 17:00 each day.
- Through the data of the epidemiological investigation, and previously conducted analyses, a plan was devised for the laboratory diagnoses of respiratory symptoms, hemorrhagic fever, and neurological symptoms. As a measure against high risk pathogens such as those causing hemorrhagic fever, the transfer of samples was determined as requiring biosafety level (BSL)-4.
- Following the transfer of the samples from the hospital on December 17th, 2019, to the BSL-4 laboratory, rapid diagnostic tests for viral hemorrhagic fevers (including Ebola and Marburg disease) were performed, and sample deactivation procedures were carried out for each case (Patients #01 and #04: blood collected right before death, and the bronchial lavage fluid, liver, and lung tissues collected after death; Patient #02: blood, throat swab, and phlegm). The rapid diagnostic test results for viral hemorrhagic fevers were negative.
- Following the transfer of the inactivated samples to a BSL-2 laboratory, DNA/RNA extraction for the genetic screening of suspected infectious diseases was carried out. The division of each task force member carried out the diagnosis by exclusion of symptoms (respiratory symptoms, hemorrhagic fever, and neurological symptoms), and the results were negative. The real-time polymerase chain reaction (PCR) or reverse transcription (RT)-PCR, and conventional PCR were performed specifically for the hemorrhagic fever virus, and possible bacteria or viruses responsible for respiratory or neurological diseases. The results were negative for hemorrhagic fever virus (including Ebola), arbovirus (including dengue fever), respiratory virus (including influenza), and malaria, Leptospira, and diphtheria.
- The cultured confluent animal cell lines (including Vero E6, MDCK, and LLC-MK2) were transferred to a BSL-4 laboratory on December 18th, 2019, and were inoculated individually with the samples from Patients #01, #02, and #04, and cultured to determine the causal pathogen. The suspected pathogenic bacteria was also cultured, and the exclusion test result indicated negative for anthrax. The second subculture of Vero E6 on December 23rd showed cytopathic effects (CPE). The receipt of samples from additional suspected individuals (friends and colleagues) and the exclusion test were carried out, and the subsequent culture test was negative for each syndrome in the exclusion list (respiratory symptoms, hemorrhagic fever, and neurological symptoms).
- The cultured cells and culture solutions were deactivated on December 23rd, 2019 and transferred to a BSL-2 laboratory where the extracted nucleic acid (DNA/RNA) underwent quality control, and libraries were prepared for next generation sequencing to be carried out. Analysis was performed on December 29th, 2019. Comparisons with national and international databases showed 97.1% sequence homology to a known coronavirus, and a request was made to the Division of Risk Assessment and International Cooperation for a literature review regarding overseas cases of novel coronavirus infections.
- The cultured cells that showed CPE were fixed and transferred to a BSL-2 laboratory on December 30th, 2019, and then an urgent request for electron microscopy analysis of the fixed cells was made to the relevant department. The analysis confirmed the presence of a coronavirus-like pathogen.
- A request for a serological test was made for serum samples from Patient #02 who was in recovery by December 23rd, 2019. The immunoserological analyses (including immunofluorescent staining, and antibody neutralization) were carried out in a BSL-4 laboratory, and immuno-serological analyses was carried out in a BSL-2 laboratory. The cultured cells clumped showing CPE, and the antibodies from the infected individual indicated that the cultured cells contained an antigen which was presumed to be the causal pathogen.
- The UDOLATF meeting was held on December 29th, 2019 for the review of results based on consultation of the relevant experts in Korea. The comprehensive analysis of results led to the confirmation of the causal pathogen as a novel coronavirus, which was reported to the director of the KCDC. On December 31st, an urgent teleconference was held to consult with overseas experts including the director-general of the WHO and the director of the CDC. These organizations provided the final confirmation of the novel coronavirus. Following the final report to the director of the KCDC, the WHO announced the emergence of a novel coronavirus and gave a press release.
- The virus was further characterized through epidemiological investigation and follow-up monitoring. No additional patient was reported for 4 weeks, a period twice as long as the incubation period after the last case, and TTX were completed with the announcement of the final case closure.
Results
2.1. Request for samples and laboratory preparation
2.2. Laboratory diagnosis and analyses
2.2.1. Diagnosis by exclusion
2.2.2. Culture of bacteria and virus
2.2.3. Next generation sequencing
2.2.4. Electron microscopy
2.2.5. Serological test
2.2.6. Review of results and confirmation of causal pathogen
2.2.7. Declaration of the end of epidemic by the novel coronavirus
- TTX were conducted for the training purposes of epidemiological investigation and laboratory response measures based on “A Guideline for Unknown Disease Outbreaks.” Considering the recent occurrences of emerging infectious diseases such as SARS and SFTS, TTX were set around a case of an unexplained respiratory disease contracted in China and imported to South Korea. The scenario was aimed at the identification of a novel coronavirus as the causal pathogen of UDO, and the performance of epidemiological investigations, management, control, and laboratory response measures.
- TTX were the first national training involving EIS officers and laboratory analysis experts regarding an outbreak of an infectious disease of unknown etiology. Through training, the members of different divisions at the KCDC could check their individual mission card, and follow the process in the identification of a novel coronavirus as the causal pathogen, based on a variety of diagnoses and analyses including the culture of bacteria and virus, next generation sequencing analysis, and molecular genetic analyses by applying the guideline-based case assessment discussion, epidemiological investigations, and algorithm per syndrome which was prepared in advance.
- For UDO, it was proposed that it is important that epidemiological investigations involve interviews, global positioning system, and credit card records to define the travel route of patients and contacts. In discussions, the participants recognized the fact that the presence of a novel coronavirus was confirmed based on pan-coronavirus RT-PCR and sequencing of SARS and MERS viruses. The use of pan-coronavirus RT-PCR was highlighted in discussions for its extensive detection of coronaviruses in unexplained coronavirus outbreaks. During the early stages of transmission when the risk of the UDO remained unclear, analyses in the BSL-4 laboratory were carried out. The possibility of diagnosis in a BSL-3 laboratory was discussed after the risk assessment for hemorrhages.
- Without prior experience of UDO, TTX reported in this study were conducted to address any weaknesses in the response process by providing an opportunity to examine and practice in advance the requirements in an actual case. The training results were effectively and appropriately utilized to respond and to identify the causal pathogen of COVID-19. Thus, TTX serve as an extremely useful practice for optimizing responses to UDO that may occur in future. Therefore, TTX should continue to be used to provide training to ensure preparedness against various emerging infectious diseases in future.
Discussion
-
Acknowledgements
- We wish to thank the KCDC EIS officers and committee members for laboratory analyses of UDO. This work was funded by the KCDC (no.: 4800-4837-301).
Acknowledgments
-
Since the government promotion on September 12th, 2020, Korea Centers for Disease Control and Prevention (KCDC) becomes Korea Disease Control and Prevention Agency (KDCA).
-
Conflicts of Interest
The authors have no conflicts of interest to declare.
Article information
- 1. Peiris JS, Lai ST, Poon LL, et al. Coronavirus as a possible cause of severe acute respiratory syndrome. Lancet 2003;361(9366). 1319−25. PMID: 10.1016/S0140-6736(03)13077-2. PMID: 12711465. PMID: 7112372.ArticlePubMedPMC
- 2. Writing Committee of the WHO Consultation on Clinical Aspects of Pandemic (H1N1) 2009 influenza. Clinical aspects of pandemic 2009 influenza A (H1N1) virus infection. N Engl J Med 2010;362(18). 1708−19. PMID: 10.1056/NEJMra1000449. PMID: 20445182.ArticlePubMed
- 3. Liu Q, He B, Huang SY, et al. Severe fever with thrombocytopenia syndrome, an emerging tick-borne zoonosis. Lancet Infect Dis 2014;14(80). 763−72. PMID: 10.1016/S1473-3099(14)70718-2. PMID: 24837566.ArticlePubMed
- 4. Zaki AM, van Boheemen S, Bestebroer TM, et al. Isolation of a novel coronavirus form a man with pneumonia in Saudi Arabia. N Engl J Med 2012;367(19). 1814−20. PMID: 10.1056/NEJMoa1211721. PMID: 23075143.ArticlePubMed
- 5. Wang ZD, Wang B, We F, et al. A new segmented virus associated with human febrile illness in China. N Engl J Med 2019;380:2116−2125. PMID: 10.1056/NEJMoa1805068. PMID: 31141633.ArticlePubMed
- 6. Anderson KG, Rambaut A, Lipkin WI, et al. The proximal origin of SARS-CoV-2. Nat Med 2020;26(4). 450−52. PMID: 10.1038/s41591-020-0820-9.ArticlePubMedPMCPDF
- 7. Centers for Disease Control and Prevention [Internet]. Unexplained respiratory disease outbreak Available from: https://www.cdc.gov/urdo.
- 8. World Health Organization [Internet]. Disease outbreak of unknown etiology Available from: https://www.who.int/environmental_health_emergencies/disease_outbreaks/unknown_etiology/en/.
- 9. Guideline for preparedness and response of unknown disease outbreak, KCDC [Internet] 2019 Available from: http://www.gbcidc.or.kr/file/download.do?file_id=277.
- 10. World Health Organization [Internet]. Guideline for the collection of clinical specimens during field investigation of outbreaks 2000 Available from: https://www.who.int/ihr/publications/WHO_CDS_CSR_EDC_2000_4/en/.
References
Figure 1Travel route of a Korean family who presented with unknown respiratory disease: Outbreak cases. A family of 4 went to Winnan, China where they hiked, visited a cave and a local farmer’s market, then took a 3-day tour of Hong Kong. The return to Korea was via Incheon on December 14th, 2019.
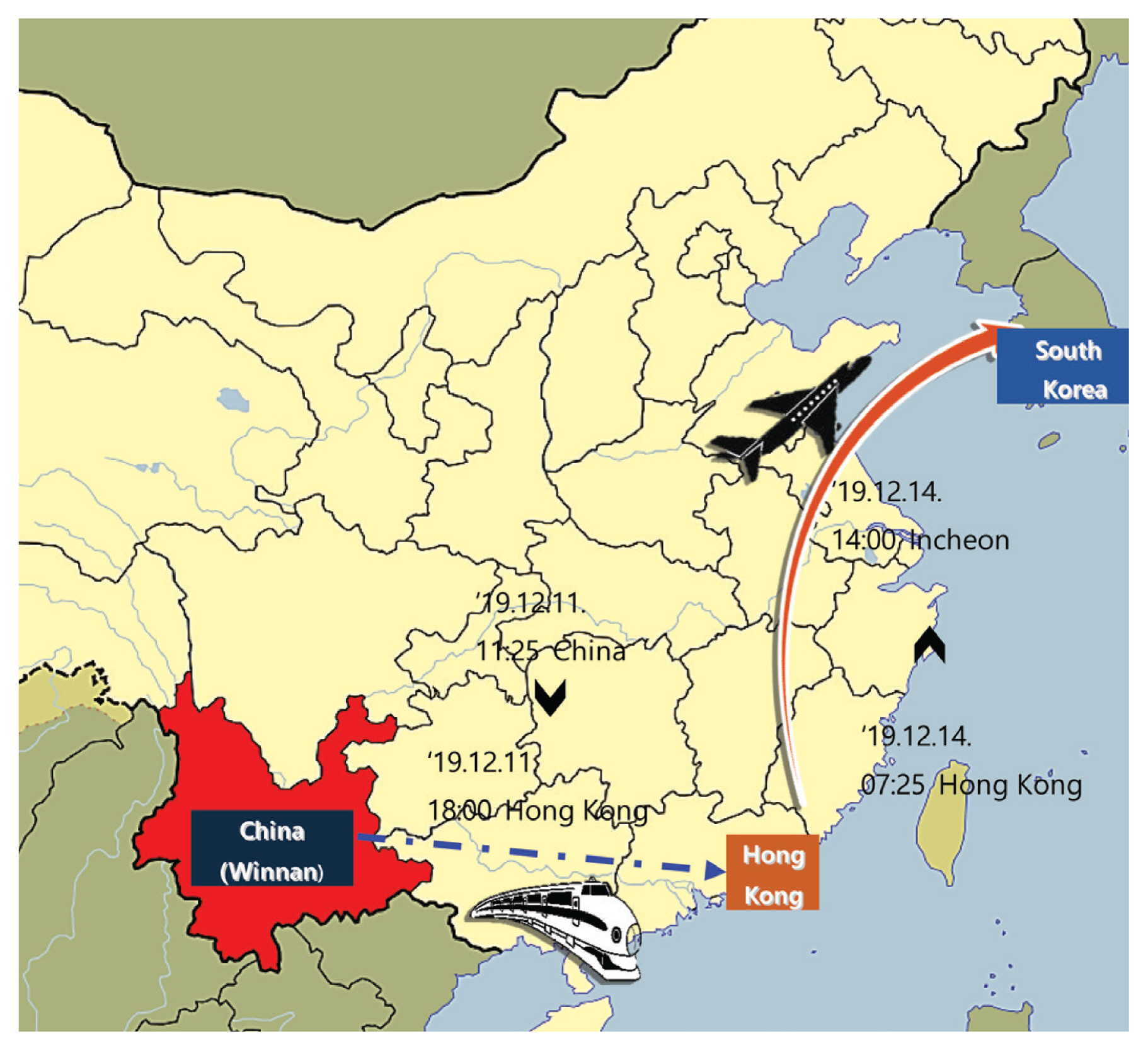

Table 1Demographics and symptoms of unknown respiratory disease outbreak.
Figure & Data
References
Citations
Citations to this article as recorded by 



 PubReader
PubReader ePub Link
ePub Link Cite
Cite