Profiling Virulence and Antimicrobial Resistance Markers of Enterovirulent Escherichia Coli from Fecal Isolates of Adult Patients with Enteric Infections in West Cameroon
Article information
Abstract
Objectives
This study aimed to identify virulent and antimicrobial resistant genes in fecal E. coli in Mbouda, Cameroon.
Methods
A total of 599 fecal samples were collected from patients with enteric infections who were ≥ 20 years old. E. coli was isolated on the MacConkey agar and virulent genes were detected by multiplex/simplex PCR. Isolates in which ≥ 1 virulent gene was detected were subjected to antibiotic susceptibility testing. The resulting resistant isolates were subjected to PCR, followed by sequencing for resistant genes detection.
Results
There were 119 enterovirulent E. coli identified, amongst which 47.05% were atypical enteropathogenic E. coli (EPEC), 36.97% enterotoxigenic E. coli, 10.08% Shiga toxin producing E. coli (STEC) and 5.88% were enteroinvasive E. coli (EIEC). The occurrence of the eae gene (47.06%) was higher compared with CVD432 (33.61%), aaic (13.45%), stx2 (10.08%) and stx1 (0.84%). High resistance rates were noted for ampicillin (94.64% EPEC, 91.67% STEC, 59.09% EAEC, and 57.14% EIEC) and sulfamethoxazole-trimethoprim (100% EPEC and 83.33% STEC, 81.82% EAEC and 71.43% EIEC). sul2 (71.43%), tetB (64.71%), tetA (59.94%) and blaTEM (52.10%) were detected. A double mutation (S83L; D87N) was seen in gyrA and a single mutation (S80I) was observed in parC.
Conclusion
These findings suggested that measures should be taken to reduce the harm of E. coli to public health.
Introduction
Escherichia coli is part of the normal enteric bacterial flora of humans and animals [1]. Nonetheless, some E. coli strains have developed pathogenic mechanisms that cause infections in humans and animals [2]. E. coli strains can be a source of enteric diarrheagenic infections in human beings [3]. The pathogenicity of E. coli is related to the association between many variables, and it is affected by ecological conditions such as microbial species, wellbeing status of the host, association with other microorganisms and antibiotic treatment received [4]. Accordingly, any E. coli conveying pathogenicity or antimicrobial resistance (AMR) is possibly pathogenic and harmful to its host [3].
Diarrheagenic E. coli (DEC) can be categorized as enteropathogenic E. coli (EPEC), enterotoxigenic E. coli (ETEC), enteroinvasive E. coli (EIEC), enteroaggregative E. coli (EAEC) or Shiga toxin producing E. coli (STEC) [5]. The categorization of E. coli as EPEC is based on the presence of either the eae gene (exterior membrane protein adhesion) and/or the bfpA gene (plasmid-encoded bundle-forming pilus) [6]. Isolated E. coli carrying the eae gene is considered as EPEC and the bfpA gene further divides EPEC (eae+) into typical and atypical [6]. E. coli is classified as ETEC based on the presence of est and elt genes which are heat-stable and heat-labile enterotoxin genes, respectively [7]. The plasmid-encoded genes CVD432 (a dispersin transporter) and aaiC (AggR-activated Island C) are used to classify EAEC E. coli isolates [8]. An enteric E. coli infection is marked by an immediate evacuation of liquid, non-bloody diarrhea of considerable volume, accompanied by little or no fever [9]. Other common symptoms are abdominal pain, malaise, nausea, and vomiting [3]. Although diarrhoeal disease is usually less harmful to adults than to children, it can reduce the productivity of its workforce which in turn affects the economy of a country. Diarrheal diseases are one of the principal causes of morbidity and mortality all over the world, especially in developing countries [10]. As reported by the Centre for Disease Control and Prevention (CDC) in 2018, diarrheal diseases are the fifth cause of deaths after HIV infection, pneumonia, lung abscess and acute bronchitis in Cameroon [11].
Bacterial diseases are treated with antimicrobial drugs [12]. Overuse of anti-infectious agents is related to the selection of AMR bacterial populations [13]. Several virulent genes and resistant genes, have been identified by PCR that code for the production of efflux pumps proteins, which decrease the permeability of the bacterial membrane, or modifying the substrate targets of antimicrobials have been identified by PCR thereby destroying antibiotic efficacy [14]. The transmission of, and pathways that may lead to, antibiotic resistance and virulence, can be divided into adaptive resistance, innate resistance, and acquired resistance. Physiological changes that lead to elevated mutation rates, i.e. mutation in metabolic genes and regulatory processes, and classic antibiotic inactivation and resistance mechanisms, could be affected by environmental factors [15]. This type of resistance and increased virulence can potentially be shared among bacteria leading to acquired resistance [15]. The rise and spread of bacterial resistance to antibiotics has been recognized as a worldwide burden, particularly in developing countries [16]. Bacterial resistance to antibiotics in the developing countries has been credited to the indiscriminate use of anti-infectious agents in animals bred for food consumption, and overuse, and imperfect prescription of antibiotics (which may influence the recovery of isolates from patients) [17].
The virulence and resistance profiles of enterovirulent E. coli strains in adults with enteric infections residing in Cameroon were determined in this study.
Material and Methods
1. Clinical sample and study population
Participants in this study were adult patients with enteric diseases, being treated in Mbouda Adlucem hospital and district hospital. These 2 health facilities are reference hospitals in the Bamboutos Division, West Region of Cameroon. Using sterile containers, a total of 599 stool specimens were sampled between May 2016 and May 2018, from patients aged ≤ 20 years. Samples were processed immediately after collection. Sociodemographic information of the study participants was collected using a structured questionnaire. Diarrheal diseases in adults are a common problem diagnosed by physicians in Mbouda. Although a diarrheal disease is usually less harmful to adults than to children, it can affect the economy of a country by reducing the output of its workforce.
2. Isolation and identification of bacteria from stool samples
Each stool sample was diluted 1:4 in sterile distilled water and inoculated onto Mac-Conkey (Becton Dickinson-Difco, Franklin Lakes, NJ, USA) agar plates. The plates were incubated in aerobiosis for 24 hours at 37°C. From morphological examination, colonies that appeared pink, flat, dry and lactose fermenting, with a bordering darker pink area of scurried bile salts were preliminarily reported as E. coli and confirmed using motility and biochemical tests such as urease production, indole production, catalase test, citrate utilization, carbohydrate fermentation test and methyl red test [18]. From each sample, 3 distinct colonies were collected for molecular determination of E. coli pathotypes (EAEC, EPEC, ETEC, EIEC and STEC) using multiplex and simplex PCR [19].
3. DNA extraction
To extract DNA, 3 distinct bacterial colonies were suspended in 200 μL of sterile phosphate buffered saline (1%). The suspension was heated to 100°C for 10 minutes and centrifuged at 10,000 g for 5 minutes. The supernatant fluid containing the DNA was stored at −20°C for supplementary molecular assays.
4. Screening of virulent genes by multiplex and simplex PCR
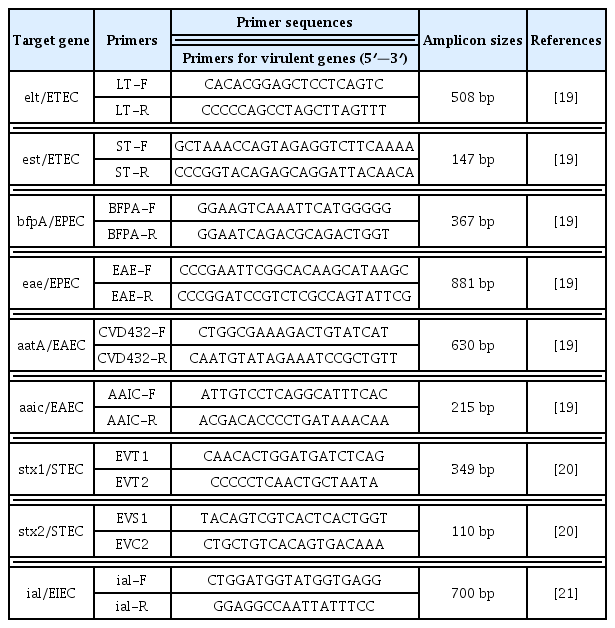
The detection of 3 types of DEC (EAEC, EPEC and ETEC) was performed using multiplex PCR testing with specific primers (Table 1) for the identification of the following virulence markers: EPEC intimin (eae gene) exterior membrane protein adhesion, EPEC plasmid-encoded bundle-forming pilus (bfpA gene), est and elt for the heat-stable and heat-labile enterotoxin genes of ETEC [7], the EAEC plasmid-encoded gene [aatA or CVD432 (a dispersin transporter)] and the EAEC chromosomal encoded aaiC locus (AggR-Activated Island C). The reaction mixture comprised of 7.4 μL of distilled water, 2.5 μL of 10x PCR buffer with 2 mM of MgCl2 (New England Biolab, UK), 0.4 μL of each LT, ST, bfpA, CVD432, aaic primer and 0.44 μL of each eae primer, 2 μL of 10 mM of dNTPs [1.25mM (New England Biolab, UK)], 0.25 μL of 5.0 U Taq DNA (New England Biolab, UK) and 3.0 μL template DNA. The PCR conditions included initial denaturation at 96°C for 4 minutes, followed by 30 cycles of denaturation at 95°C for 20 seconds, annealing at 57°C for 20 seconds and extension at 72°C for 1 minute, and a final extension at 72°C for 7 minutes. The PCR products were stained after electrophoresis on a 2% agarose gel using an ethidium bromide solution, for 25 minutes, and visualized on a transilluminator. The E. coli ATCC 29552 (EAEC), ATCC 35401 (ETEC), ATCC 43893 (EIEC), and ATCC 43895 (EPEC, EHEC, and STEC) were used as positive control strains.
All these genes are virulence determinants for their respective pathogens. Isolates positive for 1 or both of the eae and bfpA genes were assigned as EPEC. Those positive for either 1 or both EAEC factors (CVD432, aaiC genes) were designated as EAEC. All eae-positive and negative E. coli isolates from multiplex PCR were further analyzed by simplex PCR for the ial gene (invasive associated protein for EIEC detection), stx1 (shiga toxins 1) and stx2 (shiga toxins 2) virulent genes [6–8]. Isolates positive for the ial gene were assigned as EIEC, and those positive for 1 or both of the stx1, stx2 genes were considered as STEC.
5. Simplex PCR for ial gene detection
The simplex PCR reaction mixture for EIEC detection contained 15.5 μL of distilled water, 2.5 μL of 10x PCR buffer with 2 mM of MgCl2 (New England Biolab, UK), 1.0 μL of each ial primer (Table 1), 2.0 μL of 10 mM of dNTPs [1.25mM (New England Biolab, UK)], 0.20 μL of 5.0 U Taq DNA (New England Biolab, UK) and 3.0 μL template DNA. The PCR conditions consisted of an initial denaturation at 94°C for 5 minutes, followed by 30 cycles of denaturation, annealing and extension at 94°C for 30 seconds, 60°C for 30 seconds, and 72°C for 1 minute, respectively, and a final extension at 72°C for 7 minutes. Positive control strains were included in each batch of tests. The PCR products were stained after electrophoresis on a 2% agarose gel using an ethidium bromide solution, for 25 minutes, and visualized on a transilluminator.
6. Simplex PCR for stx1 (shiga toxins 1) and stx2 (shiga toxins 2) genes detection
The simplex reaction mixture comprised of 13.25 μL distilled water, 2.5 μL of 10x PCR buffer with 2 mM of MgCl2 (New England Biolab, UK), 2.0 μL of each shiga toxin primer (Table 1 [19–21]), 2 μL of 10 mM of dNTPs [1.25mM (New England Biolab, UK)], 0.25 μL of 5.0 U Taq DNA (New England Biolab, UK) and 3.0 μL template DNA. The PCR conditions involved an initial denaturation at 94°C for 5 minutes, followed by 30 cycles of denaturation at 94°C for 90 seconds, annealing at 55°C for 90 seconds, an extension at 72°C for 90 seconds, and a final extension at 72°C for 7 minutes.
7. Antimicrobial susceptibility testing
All isolates possessing at least 1 virulence gene as determined by multiplex and simplex PCR were subjected to antibiotic susceptibility testing using the Kirby Bauer disk diffusion method [22] for the following antimicrobials: ampicillin (AMP, 10μg), azithromycin (AZM, 15μg), cefotaxime (CTX, 30 μg), ceftazidime (CAZ, 30μg), ceftriaxone (CRO, 30μg), chloramphenicol (CHL, 30μg), ciprofloxacin (CIP, 5 μg), gentamicin (GEN, 10 μg), meropenem (MEM, 10μg), nalidixic acid (NA, 30 μg), norfloxacin (NOR, 10 μg), ofloxacin (OFX, 5 μg), tetracycline (TET, 5μg), and trimethoprim-sulphamethoxazole (SXT, 23.75/1.25 μg) [Becton Dickinson and Company, Sparks, MD 21152 USA]. E. coli ATCC 25922 was used as a quality control strain. The expression of the results as sensitive, intermediate and resistant was performed according to Clinical and Laboratory Standards Institute guidelines [23]. Multi drug resistance (MDR) was considered as bacterial resistance to 3 antibiotics belonging to at least 3 classes or families [24].
8. Screening of antibiotic resistant genes
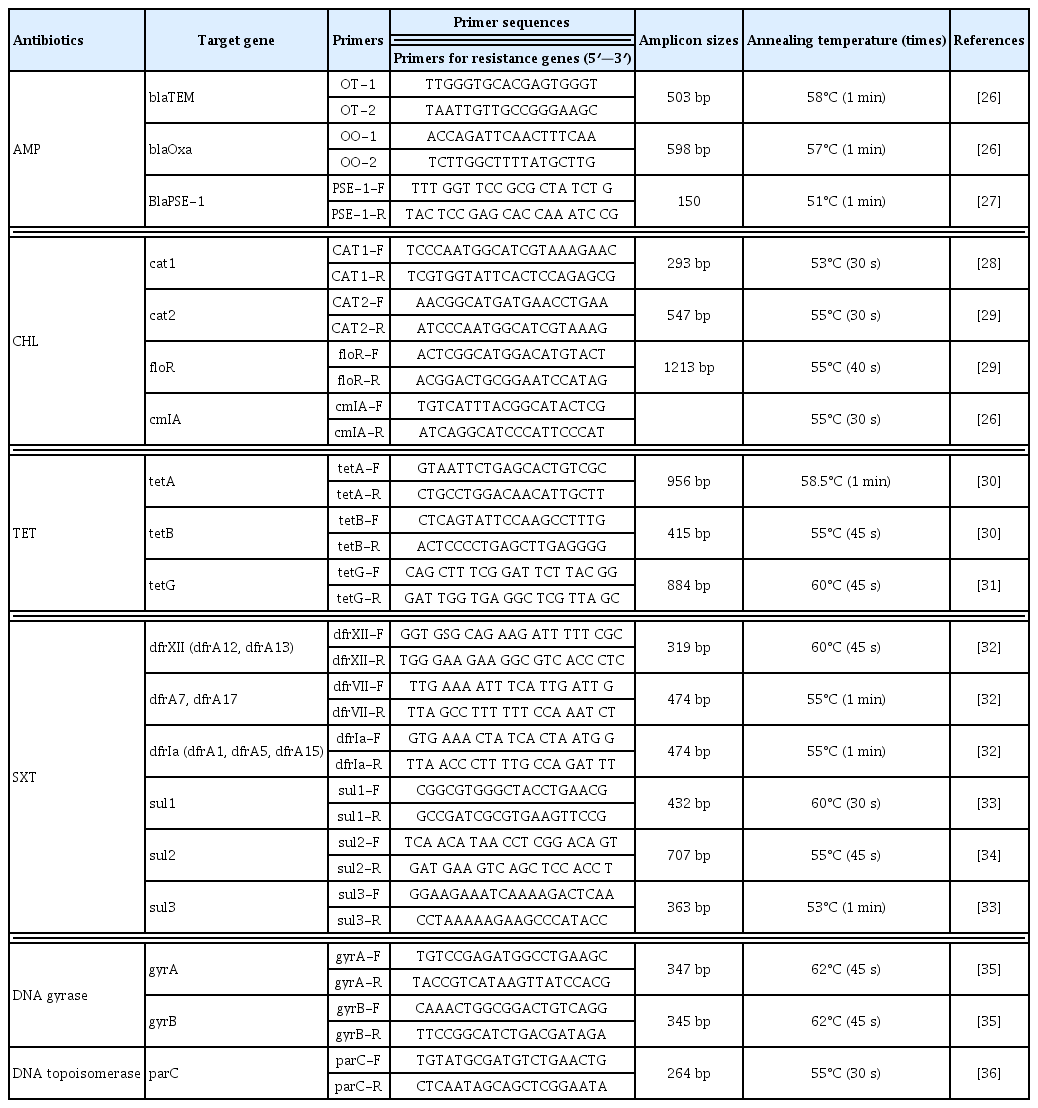
Simplex PCR was utilized for the determination of resistant genes: β-lactamases blaTEM, blaOxa and blaPSE- 1, chloramphenicol acetyltransferase (cat1 and cat2), non-enzymatic or plasmid chloramphenicol resistance (cmIA) and chloramphenicol efflux pump floR (plasmid or chromosome-encoded chloramphenicol exporter), tetracycline efflux pumps tetA, tetB and tetG, and dihydropteroate synthases sul1, sul2 and sul3 and dihydofolate reductase dfrA1, dfrA7 and dfrA12 [25]. For these antibiotic resistant genes, the PCR reaction mixture contained 8.5 μL of distilled water, 1.5 μL of 10x PCR buffer with 2 mM of MgCl2 (New England Biolab, UK), 0.15 μL of each primer, 1.5 μL dNTPs [1.25mM (New England Biolab, UK)], 0.20 μL of 5.0 U Taq DNA (New England Biolab, UK) and 3.0 μL template DNA. Table 2 shows the specific primer sequences and PCR conditions [26–36].
9. Screening of mutations in the DNA gyrase (gyrA, gyrB) and topoisomerase (parC) genes
The mutations in chromosomal DNA gyrase (gyrA, gyrB) and DNA topoisomerase (parC) carrying the quinolone resistance-determining regions were investigated using simplex PCR followed by sequencing [37,38]. For each of the gyrA, gyrB and parC genes, the PCR reaction mixture contained 8.5 μL of distilled water, 1.5 μL of 10x PCR buffer with 2 mM of MgCl2 (New England Biolab, UK), 0.15 μL of each primer, 1.5 μL dNTPs [2.5mM (New England Biolab, UK)], 0.20 μL of 5.0 U Taq DNA (New England Biolab, UK) and 3.0 μL template DNA. The PCR conditions and amplicon sizes are shown in Table 2. Amplified DNA products were resolved by electrophoresis on 2% agarose gels and stained with ethidium bromide. PCR products were purified for sequencing by BigDye XTerminator purification method (Foster City, USA). The forward and reverse strands were sequenced in Applied Biosystems Genetic Analyzers 3130/xl (Applied Biosystems) with the same PCR primer sets. Raw sequences of each gyrA, gyrB and parC genes were reviewed by visual inspection using Sequence Scanner software v2.0 (64-bit, [Applied Biosystems genetic analyzer instruments]), studied with the BLAST and FASTA programs of the National Centre of Biotechnology Information (www.ncbi.nlm.nhi.gov) and compared with the gyrA, gyrB and parC reference sequences of E. coli str. K-12 substrain MG1655 in the GenBank database.
10. Ethical considerations
Ethical endorsement was acquired from the Cameroon National Ethics Committee, Ministry of Public Health (2018/06/1054/CE/CNERSH). Before data collection, authorizations were obtained from each director of Mbouda, Adlucem and district hospitals. Information sheets presenting the objective and procedures of the study were explained to each participant. Answers were also given to their questions concerning the study. Written informed consent was obtained from each participant before his/her discretionary participation.
11. Statistical analysis
Epi Info version 7.2.2.6 (CDC, Atlanta, USA) was used for the statistical analysis. Frequencies and percentages were computed, and the study variables were compared using 2-tailed Chi-square test and correlation analysis. Results were considered significant if p < 0.05.
Results
1. Distribution of DEC pathotypes and virulent genes among total enterovirulent E. coli (n = 119)
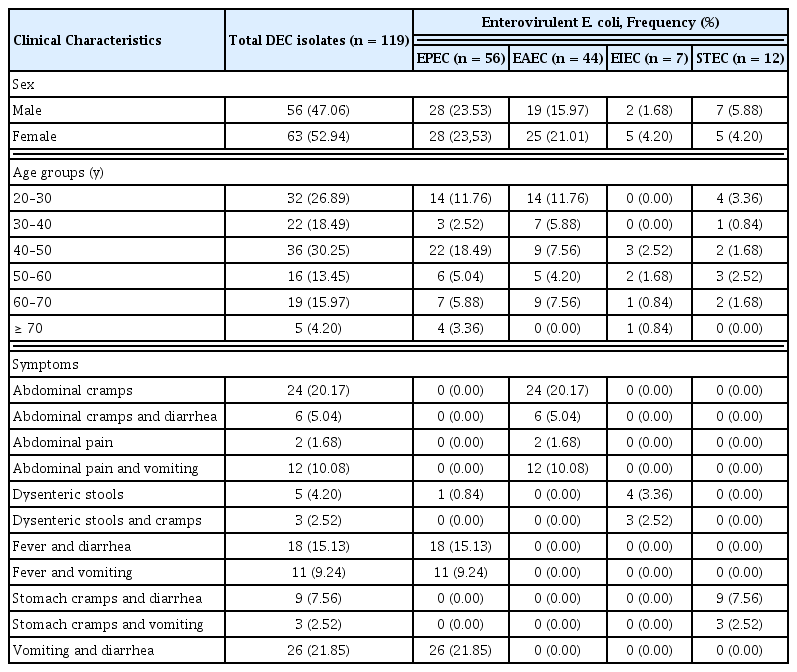
Based on macroscopic colony characteristics (pink, flat, dry and lactose fermenting colonies with a bordering darker pink area of scurried bile salts) and biochemical tests, a total of 377 E. coli isolates were obtained from 599 stool samples. Multiplex and simplex PCR used to target virulent genes enabled the identification of 119 enterovirulent E. coli isolates among which 56 (47.05%) were atypical EPEC (eae gene only), 44 (36.97%) were EAEC, 12 (10.08%) belonged to STEC, and 7 (5.88) were EIEC. None of the stool specimens was positive for typical EPEC (both eae and bfpA) and ETEC. The occurrence of DEC among females (52.94%) was slightly higher than in males (47.06%). The participant’s average age was 43.42 ± 15.61 years (ranging from 20 to 88 years) for participants in which 119 enterovirulent E. coli were detected. DEC was most prevalent in the age group 40 < 50 years (30.25%), with a predominance of EPEC (18.49%) compared with EAEC (7.56%), EIEC (2.52%) and STEC (1.68%). Vomiting and diarrhea, and fever and diarrhea, were observed in 21.85% and 15.13% of patients affected with EPEC, respectively. Patients infected with EAEC (20.17%), EIEC (3.36%) and with STEC (7.56) had abdominal cramps, dysenteric stools/stomach cramps and diarrhea, respectively (Table 3).
The molecular detection of virulence associated genes (Figure 1) showed the presence of the eae gene was predominant in 47.06% of the total isolated DEC strains, followed by the CVD432 gene (33.61%), aaic (13.45%), stx2 (10.08%) and stx1 (0.84%). The presence of associations CVD432/aaic and stx1/stx2 genes was observed in 10.08% and 0.84%, respectively, whilst the genes bfpA, and eae/bfpA were absent.
2. AMR profile of enterovirulent E. coli isolates
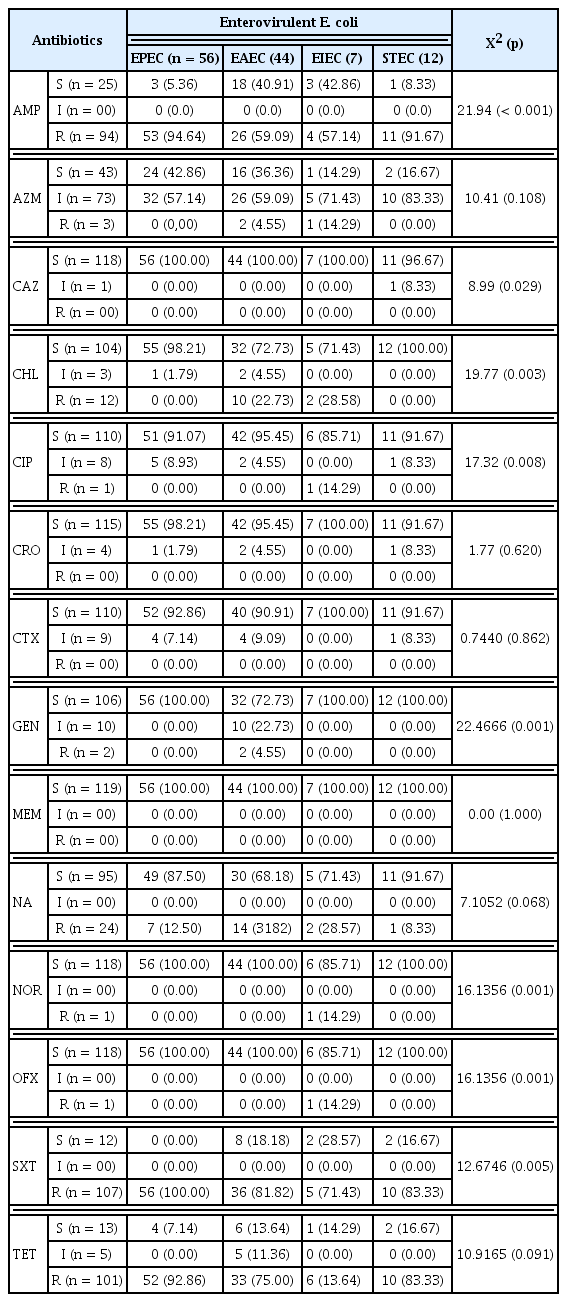
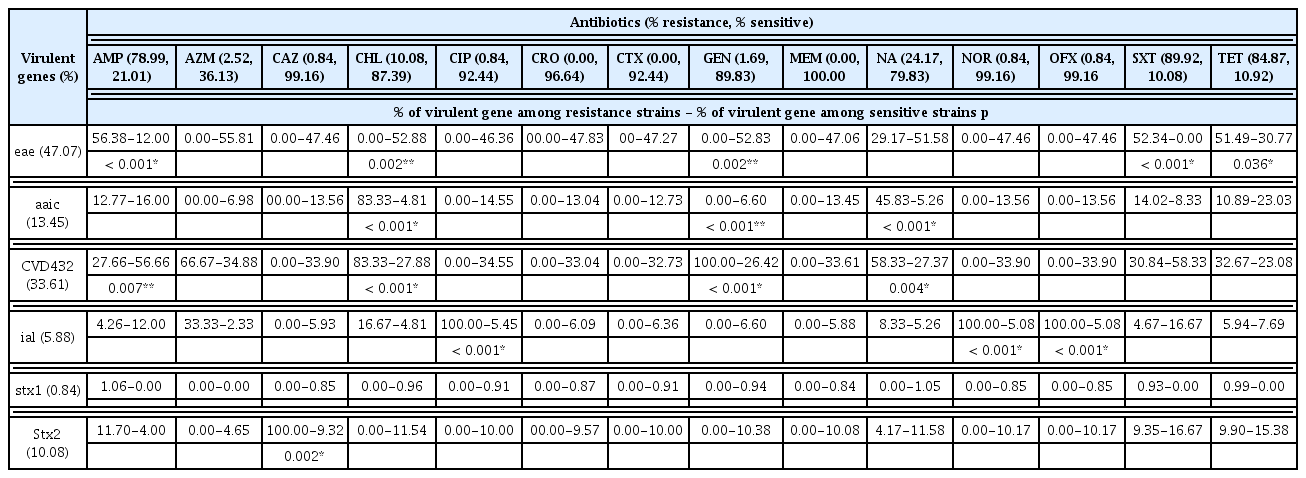
The antibiotic resistance patterns of E. coli isolates are summarized in Table 4. The E. coli isolates exhibited high resistance to antibiotics such as ampicillin (94.64% EPEC, 91.67% STEC, 59.09% EAEC, and 57.14% EIEC; X2 = 21.94, p = 0.001) and sulfamethoxazole-trimethoprim (100% EPEC and 83.33% STEC, 81.82% EAEC and 71.43% EIEC; X2 = 12.67, p = 0.005). EPEC (92.86%), STEC (83.33%), EAEC (75.00%) and EIEC (13.64) showed resistance against tetracycline although these results were not statistically significant. E. coli isolates showed significant susceptibility to the following antibiotics: ceftazidime, chloramphenicol, ciprofloxacin, gentamicin and norfloxacin. All isolates were 100% susceptible to meropenem.
3. Distribution of the antimicrobial resistant genes according to the pathotype groups
Molecular identification of resistant genes using PCR (Table 5) disclosed the existence of broad spectrum beta lactamase (BSBL) genes blaTEM (52.10%), blaOxa (9.24%) and blaPSE-1 (0.84%). The occurrence of each resistant gene was compared with the resistance profile of enterovirulent E. coli against the antibiotics. The results showed the blaTEM gene was significantly higher (p = 0.001) in ampicillin resistant STEC (75.00%), followed by EPEC (66.07%), EAEC (31.82%) and EIEC (28.57%), whilst blaOxa was significantly higher (p < 0.001) in ampicillin resistant EIEC (28.57%) compared with ampicillin resistant EAEC (20.45%), EPEC (0%) and STEC (0%). The distribution of chloramphenicol resistance genes was 10.08% for both cat1 and cat2, and 0% for both cmlA and floR. cat1 and cat2 genes were significantly higher in chloramphenicol resistant EIEC compared with other chloramphenicol resistant DEC (p < 0.001). Tetracycline resistant genes tetB, tetA and tetG, were detected in 64.71%, 59.94% and 17.65% of the total tetracycline resistant isolates, respectively. tetB was significantly higher in tetracycline resistant STEC (p = 0.006) whilst tetG was higher in EIEC (p < 0.001) compared with other tetracycline resistant DEC. Sulfonamide resistant genes sul1, sul2 and sul3 were identified in 16.81%, 71.43% and 5.88% of sulfonamide resistant isolates, respectively, and trimethoprim resistant genes dfr1A, dfrA7 and dfrA12 were present in 6.72%, 0%, and 0.84% of trimethoprim resistant isolates, respectively. sul2 was significantly higher in STEC whilst sul1 was higher in EAEC compared with other E. coli pathotypes (p < 0.05).
4. Presence of amino acid mutation in quinolone resistance-determining regions
The mutations in the chromosomal DNA gyrase (gyrA, gyrB) and DNA topoisomerase (parC) carrying the quinolone resistance-determining regions were investigated using simplex PCR followed by sequencing. Six quinolone resistant E. coli and 4 sensitive strains were sequenced. No mutation was observed in the gyrA, gyrB and parC protein sequences of all sensitive isolates. For the quinolone resistant E. coli, no mutation was observed in the gyrB protein sequence of all analyzed isolates. Alignment of the gyrA (DNA GyrAse) and parC (DNA topoisomerase) protein sequence of 1 quinolone resistant E. coli isolate is shown in Figures 2 and 3. A double mutation was observed in gyrA (accession number: MT334396) at position 83 [S83L (bestows high-level resistance)] and position 87 [D87N (bestows low-level resistance)] for a single isolate resistance to OFX, NOR, CIP and NA, whilst a single mutation in parC (accession no.: MT334398) was seen at position 80 (S80I) in the same isolate. There were 33.33% (n = 2) of E. coli isolates resistant to NA and susceptible to OFX, NOR and CIP presented a single mutation in gyrA (accession no.: MT334397) at position 83 (S83L).

Sequence alignment of the gyrA [DNA gyrase (accession no.: MT334396)] protein sequence. The substitution was seen at position 83 [S83L (confers high-level resistance)] and 87 [D87N (confers low-level resistance)]. The gyrA reference sequence was obtained from the NCBI database.
5. Associations between E. coli virulent genes and antibiotics susceptibility phenotypes
Significant associations between E. coli virulent genes and antibiotic sensitivity phenotypes were observed. Resistance to chloramphenicol was positively associated with aaic and CVD432 (p < 0.05), and negatively associated with eae (p < 0.05). Resistance to gentamicin was marked by its positive association with CVD432 and negative association with eae and aaic (p < 0.05; Table 6).
6. Associations between virulent and antibiotic resistance genes
There were significant associations between virulent genes and antibiotic resistance genes. Enterovirulent E. coli carrying blaOxa, blaTEM, cat1, cat2, sul1, and sul2 genes were distinguished by their associations. Each of the 3 virulent genes (eae, aaic, CVD432), and the cat1 and cat2 gene, were each positively associated to aaic and CVD432 gene (p < 0.05), and negatively associated to the eae gene (p < 0.05; Table 7). Concerning chloramphenicol resistance, the associations obtained genetically are in agreement with those obtained phenotypically to virulent genes.
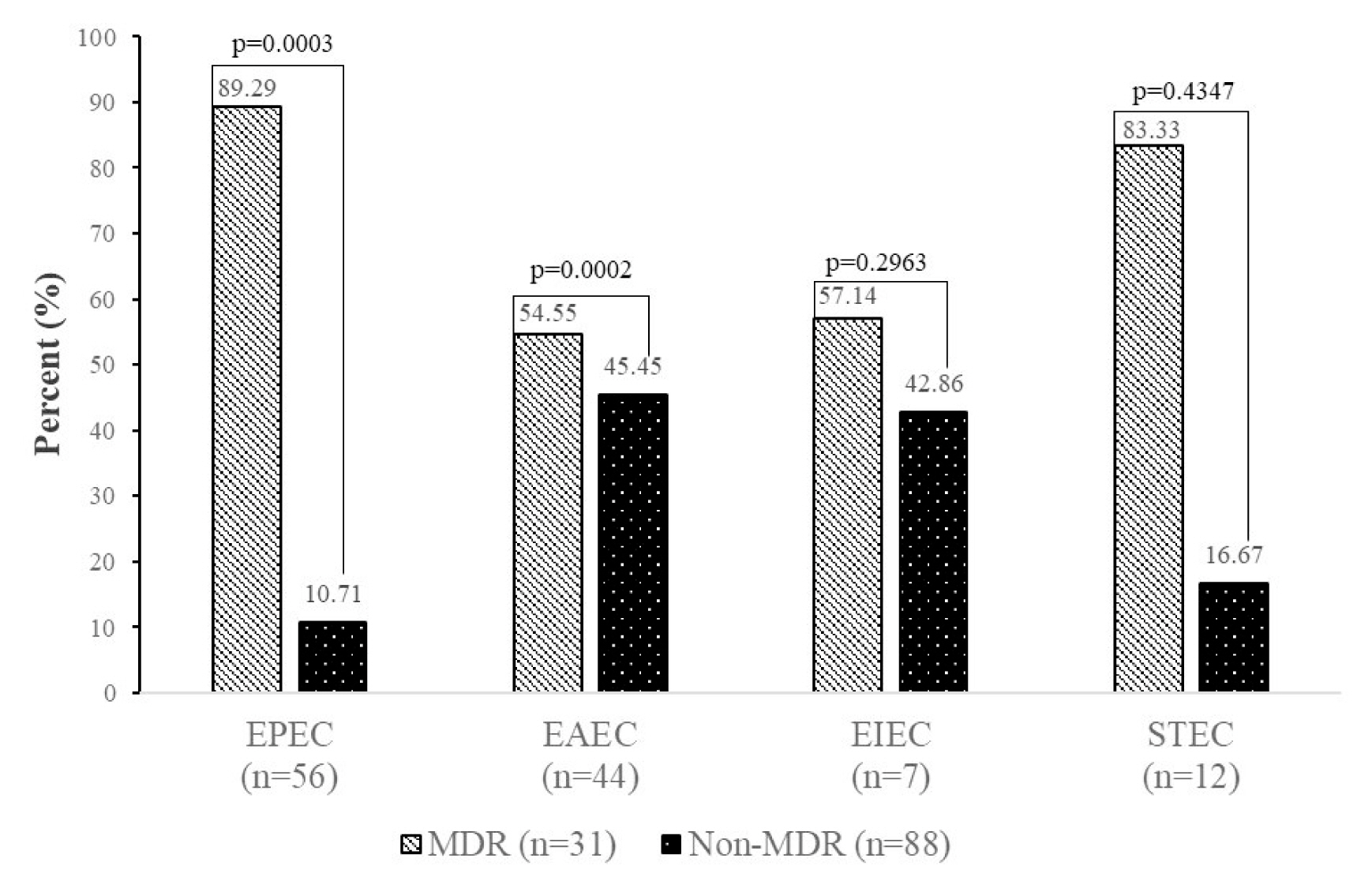
7. Multi-drugs resistant profile
The features of the E. coli pathotypes according to MDR status are presented in Figure 4. EPEC and EAEC were more likely (p < 0.05) to be MDR (89.29%, 54.55%) than non-MDR (10.71%, 45.45%), respectively.
Discussion
The diversity, as well as the pathogenicity of the E. coli strains are due to the insertion of genetic material, whether in the form of transposon, bacteriophage or plasmid [39]. This incorporated genetic material may contain genes involved in virulence and resistance against antimicrobial drugs [40]. Enterovirulent E. coli has been identified as a major cause of diarrhea in developing countries [41]. These pathogens are not routinely sought in developing countries due to deficiency of infrastructures such as advanced molecular techniques. Consequently, the precise burden of enterovirulent E. coli among adult patients throughout Cameroon, notably the Bamboutos Division, is uncertain.
In the present study, stool samples were collected from patients with enteric diseases, being treated in Mbouda, Adlucem and district hospitals, in the Bamboutos Division, West region of Cameroon. This is the first in-depth study of enterovirulent E. coli in the population of Mbouda. The occurrence of atypical EPEC was higher compared with EAEC, STEC and EIEC. None of the stool specimens were positive for typical EPEC (both eae and bfpA) and ETEC. Similar results obtained in Mangalore, India showed that, the most prevalent DEC was atypical EPEC, followed by EAEC and STEC [42], contrary to that of Spano, Brazil [43].
The occurrence of EPEC was higher in patients with symptoms of vomiting and diarrhea, and fever and diarrhea, while patients infected with EAEC, EIEC and STEC reported abdominal cramps, dysenteric stools/stomach cramps and diarrhea, respectively. It has been shown that, various E. coli pathotypes contribute to diarrheal episodes by inhibiting the serotonin transporter, although the mechanism is poorly understood [6]. Among all enterovirulent E. coli, EPEC was observed to be the most widespread pathotype for diarrheal infection in Mbouda.
The molecular detection of virulent associated genes revealed that, the occurrence of the eae gene in total screened isolates was high compared with CVD432, aaic, stx1 and stx2. The association of the CVD432/aaic and stx1/stx2 genes was observed in 10.08% and 0.84%, respectively, whilst the other genes (bfpA, eae/bfpA) were absent. These findings are comparable to those obtained in a study reported in India [42]. The distribution of E. coli pathotypes and virulent genes depends on geographical and socio-economic features of each region. In Tehran, Shahrokhi et al showed that, ST was the most frequent toxin type followed by LT and LT/ST [44]. Dominance of the ST-expressing ETEC has been documented in Egypt, Bangladesh, and Iran [44,45].
Highly significant resistance rates were observed for ampicillin and sulfamethoxazole-trimethoprim in this study. EPEC (92.86%), STEC (83.33%), EAEC (75.00%) and EIEC (13.64) showed non-significant resistance to tetracycline. These results are comparable to the global general trend of fecal E. coli isolates [3]. Nevertheless, resistance rates fluctuate by country. In comparison, the rates observed in this study for Cameroon are higher than those registered in 6 European countries including Russia [46], and lower than those from Nigeria [47]. The spectrum of antibiotics used in Cameroon for prophylaxis, therapy, growth promotion and auto-medication can directly affect AMR of endogenous bacteria. In addition, the environment can also be a source for resistant microorganisms and resistant genes for humans [48].
BSBL genes blaTEM and blaOxa, were detected in a wide number of isolates, contrary to an existing report [49]. This may have been the cause of resistance to ampicillin in the study isolates. The prevalence of chloramphenicol resistant genes cat1 and cat2, was higher in E. coli isolates resistant to chloramphenicol compared with the cmlA and floR genes. This indicates non-enzymatic or plasmid chloramphenicol resistant mechanisms of study isolates [50]. Tetracycline genes tetB, tetA, and tetG, were detected in 64.71%, 59.94% and 17.65% of the total tetracycline resistant isolates, respectively, with a predominance among tetracycline resistant STEC for tetB and EIEC for tetG. These results show that, study isolates express resistance via an efflux mechanism [51]. sul1, sul2 and sul3 (dihydropteroate synthase genes) genes were detected in 16.81%, 71.43% and 5.88% of trimethoprim-sulphamethoxazole resistant isolates, respectively. These genes play a major role in sulfonamide resistance and are remarkably related to integrons and transposons. High occurrence of the sul2 gene observed in this study is in accordance with the results obtained by Wu et al in 2010 [52]. The genes dfr1A, dfrA7 and dfrA12 were detected in 6.72%, 0.00%, and 0.84% of trimethoprim resistant isolates, respectively, dfr1A was the most commonly identified. The occurrence of antibiotic resistance fluctuates with respect to each country. These findings are similar to those from a report in Iran [53].
A double substitution (S83L; D87N) was seen in the gyrA protein sequence for a single isolate resistant to OFX, NOR, CIP and NA whilst a single mutation in parC was seen at position 80 (S80I). 33.33% E. coli isolates resistant to NA and susceptible to OFX, NOR and CIP presented a single mutation in gyrA at position 83 (S83L). Many factors such as animal-to-human transfer of resistant bacteria have been reported to be the main contributor to the appearance of highly quinolone-resistant bacteria, mainly due to mutations in the DNA gyrase and topoisomerase IV genes [54].
Many associations between virulent factors and AMR were identified. The most remarkable was chloramphenicol resistance that was positively associated with aaic and CVD432, and negatively associated with eae, and that of gentamicin which was distinguished by its positive association with CVD432 and negative association with eae and aaic. The combination of E. coli virulent factors and antibiotic resistance has been reported although the molecular pathways underlying the association between resistance and virulence is not clearly understood.
Other associations were observed between virulent genes and antimicrobial resistant genes. Enterovirulent E. coli carrying the blaOxa, blaTEM, cat1, cat2, sul1, and sul2 genes, were distinguished by their association with each of the 3 virulent genes (eae, aaic, CVD432). cat1 and cat2 genes were each positively associated with aaic and the CVD432 genes, and negatively associated with the eae gene. Concerning chloramphenicol resistance, the associations obtained genetically are in agreement with those obtained phenotypically to virulent genes.
These findings are of huge public health prominence since the virulent and resistant genes of enterovirulent E. coli are not routinely assessed in hospital-based laboratories in Mbouda, Cameroon. However, the study population in this present study was limited to patients aged ≥ 20 years. In addition, the focus of this work was on enterovirulent E. coli isolates. Even though non-enterovirulent E. coli isolates were obtained, no further molecular studies were performed. Enterovirulent E. coli is reputed to carry resistant genes on plasmids, transposons, and integrons. These mobile genetic materials can be transferred between microorganisms of the same species or different genera across horizontal gene transfer [40]. Thus, additional studies are needed to evaluate the risks associated with E. coli which carry antibiotic resistance genes, and the capability of transferring these genes to other bacteria, including commensal E. coli and other enteric bacterial pathogens which could have a prejudicial effect on public health.
Conclusion
This study, the first in Mbouda, Cameroon to determine the virulence and antibiotic resistance of enterovirulent E. coli isolates from a sample of the Mbouda adult population with enteric diseases. The presence of atypical EPEC, EAEC, STEC, and EIEC was observed. A high antibiotic resistance phenotype was noted, with resistance to commonly used antibiotics including ampicillin, sulfamethoxazole-trimethoprim, and tetracycline. Significant susceptibility to antibiotics such as ceftazidime, chloramphenicol, ciprofloxacin, gentamicin, norfloxacin and meropenem, was equally observed. BSBL genes blaTEM and blaOxa, chloramphenicol resistant genes (cat1 and cat2), tetracycline resistant genes (tetB, tetA and tetG), dihydropteroate synthase genes (sul1, sul2 and sul3) and trimethoprim resistant genes (dfr1A, dfrA7 and dfrA12) were detected in the respective antibiotic resistant enterovirulent E. coli. Many associations between virulent factors and antimicrobial resistant phenotype, and between virulent genes and antimicrobial resistant genes, were noted. The detection of virulent and highly antibiotic resistant genes in fecal E. coli in Mbouda adults suggested that measures should be taken to reduce the potential harm to public health, such as the appropriate etiological diagnosis and reasonable use of antimicrobial agents.
Appendix
Appendix
Supporting files are available at https://www.kcdcphrp.org.
Supporting file1 (.xls): Raw data of the study.
Supporting file2 (.doc): Figure S1. Nucleotide sequence of the gyrA DNA sequence (accession number: MT334396) form sample 60J whose a Compositional matrix adjust analyses show A double Mutation in gyrA protein sequence at position 83 (S change to L= S83L) and 87 (D change to N= D87N);Figure S2. Nucleotide sequence of the gyrA DNA (accession number: MT334398) sequence form sample 90J3 whose a Compositional matrix adjust analyses show A single Mutation in gyrA protein sequence at position 83 (S change to L= S83L); Figure S3. Nucleotide sequence of the parC DNA sequence (accession number: MT334397) fragment form sample 60J whose a Compositional matrix adjust analyses show A single Mutation in parC protein sequence at position 80 (S change to I= S80I).
1. Data availability
All data generated or analyzed during this study are included in this published article and the supporting file. Sequences have been deposited in NCBI GenBank under the accession numbers MT334396; MT334398 and MT334397.
Acknowledgments
Mr. Marbou wishes to thank the World Academy of Sciences (TWAS) and Department of Science and Technology (DBT) for his fellowship (Award of 2017 DBT-TWAS PG, RF number: 3240300012) and Dr. Shanta Dutta, the director of the National Institute of Cholera and Enteric Diseases for accepting him as fellow at the Indian Council of Medical Research-National Institute of Cholera and Enteric Diseases, Kolkata, India. Mr. Marbou acknowledges the working facilities of Division of Bacteriology, and all the staff, technicians, research fellows for their un-stint support. He is also thankful to Mr. Karmakar for his cooperation.
Notes
Conflicts of Interest
The authors have no conflicts of Interest to declare.