Articles
- Page Path
- HOME > Osong Public Health Res Perspect > Volume 1(1); 2010 > Article
-
Original Article
Distribution of Virulence Genes and Their Association of Serotypes in PathogenicEscherichia coli Isolates From Diarrheal Patients in Korea - Seung-Hak Cho, Kyung-Hwan Oh, Seong-Han Kim, Hee-Bok Oh, Mi-Sun Park
-
Osong Public Health and Research Perspectives 2010;1(1):29-35.
DOI: https://doi.org/10.1016/j.phrp.2010.12.008
Published online: December 7, 2010
Division of Enteric Bacterial Infections, Center for Infectious Diseases, Korean National Institute of Health, Korea Centers for Disease Control & Prevention, Korea
- ∗Corresponding author. Division of Enteric Bacterial Infections, Center for Infectious Diseases, Korean National Institute of Health, Korea Centers for Disease Control & Prevention, Nokbeon-dong 5, Seoul 122-701, Republic of Korea. pmsun63@korea.kr
© 2010 Published by Elsevier B.V. on behalf of Korea Centers for Disease Control and Prevention.
This is an Open Access article distributed under the terms of the Creative Commons Attribution Non-Commercial License (http://creativecommons.org/licenses/by-nc/3.0) which permits unrestricted non-commercial use, distribution, and reproduction in any medium, provided the original work is properly cited.
Abstract
-
Objectives
- To characterise the genetic and serological diversity of pathogenic Escherichia coli, we tested 111 E coli strains isolated from diarrhoeal patients in Korea between 2003 and 2006.
-
Methods
- The isolates were tested through polymerase chain reaction (PCR) and slide agglutination method for the detection of virulence genes and serotypes, respectively. To compare the expression of Shiga toxin (stx)-1 and stx2 genes, real-time quantitative reverse-transcriptase PCR and rapid exprssion assay, reversed-passive latex agglutination, were performed.
-
Results
- Forty-nine Shiga toxin-producing E coli (STEC) strains and 62 non-STEC strains, including 20 enteropathogenic E coli, 20 enterotoxigenic E coli, 20 enteroaggregative E coli, and 2 enteroinvasive E coli were randomly chosen from the strains isolated from diarrhoeal patients in Korea between 2003 and 2006. PCR analysis indicated that locus of enterocyte effacement pathogenicity island, that is, eaeA, espADB, and tir genes were present in STEC, enteropathogenic E coli, and enteroinvasive E coli. Quorum sensing-related gene luxS was detected in most of pathogenic E coli strains. Major serotypes of the STEC strains were O157 (26%) and O26 (20%), whereas the non-STEC strains possessed various serotypes. Especially, all the strains with serotype O157 carried stx2 and the tested virulence factors. Of the STEC strains, the data of real-time quantitative reverse-transcriptase PCR and reversed-passive latex agglutination tests showed that messenger RNA- and protein expression of stx2 gene were higher than those of stx1 gene.
-
Conclusion
- Our results provide the epidemiological information regarding the trend of STEC and non-STEC infections in the general population and show the fundamental data in association of serotypes with virulence genes in diarrhoeagenic E coli strains from Korea.
- Diarrhoea is an extraordinarily common disease with worldwide distribution, and diarrhoeagenic Escherichia coli is an important bacterium to cause diarrhoeal disease.1,2 In a surveillance of bacterial pathogens associated with acute diarrhoeal disease in the Republic of Korea, it has been found that pathogenic E coli are frequently isolated from diarrhoeal patients (around 20%), and enterohaemorrhagic E coli (EHEC) accounts for ca. 2% among the isolated pathogenic E coli.3 The pathogenic strategies of the diarrhoeagenic E coli strains exhibit remarkable variety. Three general paradigms have been described by which E coli may cause diarrhoea;2 each is described in detail in the appropriate section below: enterotoxin production [enterotoxigenic E coli (ETEC) and enteroaggregative E coli, (EAEC)], invasion [enteroinvasive E coli (EIEC)], and intimate adherence with membrane signalling [enteropathogenic E coli (EPEC) and EHEC].4–7
- The major virulence factor, which is a defining characteristic of EHEC, is Shiga toxin (Stx). Shiga toxin-producing E coli (STEC) strains produce one or both of two major types of shiga toxin, designated Stx1 and Stx2, and the production of the latter is associated with an increased risk of developing haemolytic-uremic syndrome.1,8 The prototypical Stx1 and Stx2 toxins have 55% and 57% sequence homology in the A and B subunits, respectively.2,6 ETEC strains are identified by the ability to produce enterotoxins, heat-labile toxin (LT), heat-stable toxin (ST), and surface adhesions known as colonization factors.7 Atypical EPEC strains would possess the attaching and effacing (eae) gene that correlates with possession of 35-kb locus of enterocyte effacement (LEE) pathogenicity island encoding eae.9 LEE encodes a Type III secretion system and E coli secreted proteins through Type III secretion system, which deliver effector molecules to the host cell and disrupt the host cytoskeleton.4,5,10,11 LEE also carries eae, which encodes an outer membrane protein (intimin) required for intimate attachment to epithelial cells; therefore, eae has been used as a convenient diagnostic marker for LEE-positive EPEC strains.9 EAEC strains produce enteroaggregative heat-stable enterotoxin (east)-1 encoded by east1 gene.12,13 Furthermore, other virulence factors of pathogenic E coli for the diarrhoeagenic infections, such as EHEC hemolysin (E-hly), espADB (Type III secretion proteins), and tir (translocated intimin receptor), have been found in diarrhoeagenic E coli strains. Quorum sensing is a mechanism of cell-to-cell signalling involving the production of hormone-like compounds called autoinducers.14,15 Regulation of LEE genes by quorum sensing is reported.16 When these molecules reach a certain concentration threshold, they interact with bacterial regulatory proteins, thereby controlling gene regulation mechanism in both gram-negative and gram-positive bacteria.14,15 The gene encoding the autoinducer-2 synthetase was cloned, sequenced, and named luxS. Recently, the global regulation of virulence factors was investigated in clinical EHEC isolate in Korea by creating a ΔluxS mutant strain.17
- To obtain the epidemiological information on various virulence factors in different pathogenic E coli groups, we characterized genetic diversity of E coli strains isolated from diarrhoeal patients in Korea from 2003 to 2006 through the surveillance system performed by Laboratory of Enteric Infections in Korean National Institute of Health.
Introduction
- 2.1 E coli strains isolated from stool specimen
- A total of 111 E coli strains isolated from the stool of diarrhoeal patients in Korea between 2003 and 2006 were investigated in this study. Bacteria were plated on MacConkey agar. All isolates were biochemically identified with the API20E system (bioMérieux, Marcy l’Etoile, France).
- 2.2 Detection of virulence genes by PCR
- Bacteria was directly inoculated into 3 mL of Luria-Bertani broth for enrichment and incubated overnight at 37°C under shaking conditions. After incubation, enriched broth culture was centrifuged at 13,000 rpm (Sorvall® Biofuge Pico, Germany) for 1 minute, and the pellet was heated at 100°C for 10 minutes. After centrifugation of the lysate, 5 μL of the supernatant was used in the polymerase chain reactions (PCRs). To identify virulence genes, PCR assays were performed using in the primers shown in Table 1. PCR assays were carried out in a 50 μL volume with 2U DNA Taq polymerase (Takara Ex Taq™, Japan) in a thermal cycler (PTC-100; MJ Research, Watertown, MA, USA) under the following conditions: initial denaturation at 94°C for 5 minutes, 30 cycles of 94°C for 1 minute, 72°C for 1 minute, and final cycle 72°C for 5 minutes. Amplified PCR products were analysed by gel electrophoresis in 2% agarose gels stained with ethidium bromide, visualized with ultraviolet illumination, and imaged with the Gel Doc 2000 documentation system (Bio-Rad, Hercules, CA, USA).
- 2.3 Total RNA isolation
- RNA extracts of the strains were prepared using the Qiagen RNeasy midi-prep kit and RNA BacteriaProtect (QIAGEN Co. Ltd, Germany) according to the manufacturer’s instructions for gram-negative bacteria. 2.3.1
- For quantitative real-time reverse-transcriptase-PCR (qPCR), the 20 μL reaction mixture was prepared by 2 μL of total RNA, 0.6 μM of each primer, and reference dye Synergy Brands Inc. Green. The number of copies was calculated, and dilutions ranging from 100 pg to 100 ng copies of this standard were prepared in a Tris-ethylenediaminetetraacetic acid buffer. Aliquots of these dilutions were frozen at −20°C. Throughout this study, the Quantitect Synergy Brands Inc. Green master mix kit (Qiagen, Valencia, CA, USA) was used for all reactions with qPCR. The parameters for qPCR included 30 minutes incubation at 50°C for converting messenger RNA (mRNA) to complementary DNA. Subsequent amplification of complementary DNA was carried out by using an initial cycle of 95°C for 10 minutes followed by 40 cycles of 94°C for 30 seconds, 55°C for 30 seconds, and 72°C for 60 seconds. The final extension was carried out at 72°C for 2 minutes. For the quantification, of stx1, stx2, and gapA (housekeeping gene) mRNA, a negative control, consisting of nuclease-free water in place of template DNA, was included in each PCR run. The primers used for real-time PCR were F 5′-GATGATCTCAGTGGGCGTTC-3′, R 5′- CAACTCGCGATGCATGATG-3′ (stx1), F 5′-CGCACTGTCTGAAACTGCTC-3′, R 5′- TCGCCAGTTATCTGACATTC-3 ′(stx2), F 5′-ACTTCGACAAATATGCTGGC-3′, R 5′- CGGGATGATGTTCTGGGAA-3′ (gapA). Reaction conditions for amplification and parameters for fluorescence data collection were programmed into an Opticon Monitor Software package 1.4 (DNA engine Opticon 2; MJ Research, Watertown, MA, USA). All assays were performed in triplicate. The threshold cycle (Ct) values of the known standards were plotted versus the logarithm of the concentration of each standard creating a standard curve. Samples of unknown concentration were plotted onto the standard curve to calculate their concentration. Normalization of the quantification results from stx1 and stx2 was performed by the incorporation of the quantification results of gapA mRNA into the following equation:Here, ΔCt endogenous gene (control − test) = Ct value of the endogenous gene (gapA) with the control RNA − Ct value of the endogenous gene (gapA) from the sample RNA. Also, ΔCt target gene (control − test) = Ct value of the target gene (stx1 or stx2) with the control RNA − Ct value of the target gene (stx1 or stx2) with the test RNA.
- 2.4 Reversed-passive latex agglutination test for the detection of Shiga toxin
- The production of Stx1 and Stx2 by the isolates was determined by using a reversed-passive latex agglutination (RPLA) kit (VTEC-RPLA; Denka Seiken Co., Ltd., Tokyo, Japan) after having been grown and shaken in 5 mL of Tryptone Soya Broth overnight at 37°C. Of this suspension, 1 mL was centrifuged for 20 minutes at 13,000 rpm. The titre of the supernatant was determined in the veterotoxin-producing E coli-RPLA test according to the manufacture’s instructions up to 1:256. All STEC strains were tested for the production of Stx1 and Stx2. Titres lower than 1:2 were interpreted as a negative control.
- 2.5 Serotyping of O antigen
- The presence of O antigens was determined by slide agglutination with the method of Guinée et al.18 using all available O antisera (O1–O181). All antisera were absorbed with the corresponding cross-reacting antigens to remove the non-specific agglutinins.
Materials and Methods
2.3.1 Amplification of target genes by real-time reverse-transcriptase-PCR and analysis
- 3.1 Diversity of virulence genes in pathogenic E coli isolates from diarrhoeal patients
- To characterize genetic diversity of virulence factors, 111 pathogenic E coli strains, 49 STEC strains, and 62 non-STEC strains, were chosen from the strains isolated from diarrhoeal patients in Korea between 2003 and 2006. All the strains were tested through PCR method with the primers as shown in Table 1. As shown in Table 2, in the genomic DNA of the 49 STEC strains, 22 strains were stx1 positive and 17 were stx2 positive. Ten possessed both stx1 and stx2 genes.
- To detect major virulence genes for pathogenic E coli, eaeA gene, major virulence factor of EPEC, lt and st genes for ETEC, east1 gene for EAEC, and invasion-associated locus gene for EIEC were analysed in the STEC and non-STEC strains by PCR with the specific primers for these genes. Among the non-STEC strains, 20 EPEC, 20 ETEC, 20 EAEC, and 2 EIEC strains were randomly chosen among the pathogenic E coli strains isolated from diarrhoeal patients in Korea. The eaeA gene was present in EPEC, STEC, and EIEC strains. The lt and st genes showed different prevalence in the pathogenic E coli strains. The lt gene was found in various pathogenic E coli groups: ETEC, STEC, EPEC, and EIEC; whereas the st gene was detected only in ETEC strains. Interestingly, the east1 gene was also distributed in various pathogenic E coli groups except EIEC strains. Eighty-five percent (17 of 20 strains) of ETEC strains harboured east1 gene. This gene was detected in 16% of STEC and 25% of EPEC strains. However, invasion-associated locus gene was found only in EIEC strains (Table 2).
- The prevalence of hlyA, espADB, tir, and luxS genes in STEC and non-STEC strains was determined in pathogenic E coli isolates. Seventy-one percent (35 of 49 strains) of the STEC strains harboured the hlyA gene, whereas all the ETEC, EAEC, and EIEC strains were negative for this gene. Among the EPEC strains, only 10% were positive for this gene. LEE-associated genes, espADB, and tir genes were present in 61%–67% of STEC strains: espA (63%), espD (67%), espB (61%), and tir (60%). Interestingly, espA and tir genes in the EPEC strains were less prevalent than espD and espB genes. All EPEC strains harboured espD and espB genes, whereas espA and tir genes were present only in 55% of the strains. Among the ETEC and EAEC strains, LEE-associated genes were not detected. These genes were found in the one of the two EIEC strains. Most of the pathogenic E coli strains carried luxS gene. This gene was present in all ETEC strains and 97% of the STEC strains. The presence of this gene was similar in EPEC (55%), EAEC (70%), and EIEC (50%) strains (Table 2).
- 3.2 Serotyping of pathogenic E coli isolates from diarrhoeal patients
- As shown in Table 3, the O antigen was typeable for 95 strains, which could be classified into 38 different serotypes, and 15 strains were non-typeable. Among the STEC strains, 12 serotypes were identified. Major serotypes of the STEC strains were O157 (26%) and O26 (20%). Under the non-STEC strains, the EPEC, ETEC, and EAEC strains showed diversity of serotypes. However, the serotype of two EIEC strains was non-typeable.
- 3.3 Association of serotypes of pathogenic E coli isolates with virulence genes
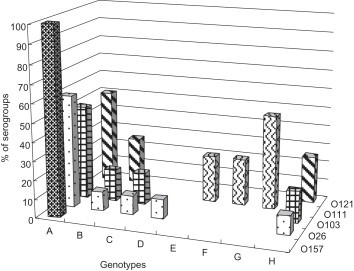
- The serotypes of STEC strains were examined in association with virulence genes. The data of serotypes in association with stx genes were described in Table 4. The stx1 gene was mostly distributed between the O26 (80%) and O103 (100%) serotypes. The stx2 gene was present in the great majority of the O157 (100%) and O121 (100%) serotypes. As shown in Figure 1, all O157 strains harboured all the tested genes, that is, eaeA, hlyA, espADB, and tir genes. The serotypes that possessed all the genes described above were O26 (6 of 10 strains), O103 (three of six strains), and O121 (two of four strains), except for serotype O157 (Group A). Among the four O26 strains, eaeA-negative strain (Group B), hlyA-negative strain (Group B), espD-negative strain (Group C), and eaeA, hlyA, espADB, tir-negative strain (Group H) were found.
- 3.4 Expression of stx genes in STEC O157 strains
- To compare mRNA and protein expression of stx1 and stx2 genes, qPCR and rapid expression assay, RPLA were performed in the STEC O157 strains (n = 13). Seven strains possessed stx2 gene only and six strains carried both stx1 and stx2 genes. For each isolate, analysis of gene expression of gapA, stx1, and stx2 was performed in the same PCR run. Application of qPCR assays indicated that the mRNA expressions of stx2 gene were higher than those of stx1 gene. Using dilution gradients of culture supernatant fluids by RPLA, expression titres of stx genes were given. The expression titres of Stx2 was also higher than those of Stx1 (Table 5). However, in some strains, low expressions of stx2 gene were found; for example, mRNA expression of Strain 2 and protein expressions of Strains 3 and 4.
Results
- The present study employed a range of several E coli organisms in the aetiology of diarrhoeal patients in Korea providing to the genetic characterization with regard to their harbouring of potential virulence genes.
- To obtain the genetic and serological diversity of pathogenic E coli, the association between the virulence factors and serotypes of isolates found in human was examined in this study. We showed that the genotypes of O157 serotype and non-O157 serotypes in STEC strains were different. The results indicated that all O157 serotypes of the STEC strains carried all the tested virulence genes, whereas these genes were detected in a lesser percent of the non-O157 STEC strains. Moreover, the stx2 gene was present in the great majority of the O157. These crude data suggest an association of stx2 with isolates of serotypes found in humans with severity of disease. O’brien et al.19 reported that 67 EHEC O157 strains tested possessed the EHEC hlyA gene. Boerlin et al.20 reported that eaeA and stx2 were significantly more frequent in isolates from serotypes found in humans with severe disease. It has been shown that the chromosomal virulence genes of EHEC and EPEC are organized as a cluster referred to as a pathogenicity island.5 Our data showed that the genotype in STEC and EPEC was similar (Table 2). Genes for both classes are found predominantly on plasmids, and some ST-encoding genes have been found on transposon.2 STa has about 50% identity to east1 of EAEC. It has recently been reported that some strains of ETEC may also express east1 in addition to STa.2,12 STb has been found only in ETEC.2 The main conclusion of these previous investigations is that no single factor is responsible for the virulence of E coli strains.
- The second part of our study showed differences of stx genes in O157 strains that are known as a pathogen in association with severe disease. Previous studies have shown that the virulence of STEC for humans may be related to the type of stx, which is produced by the bacteria and serotype.21 In a study concerning Stx production as a single microbial factor, the most pathogenic strains for humans have been found to produce Stx2 only.2 The Stx2 toxin has been described as being 1,000 times more cytotoxic than Stx1 towards human renal microvascular endothelial cells.2,8 In other studies, Stx2 was found to be related with high virulence and was significantly associated with STEC strains from haemolytic-uremic syndrome patients.8 Our data indicated that the expression of stx genes was different. Although low expressions of stx2 gene were found in some strains (Table 5), we suggest that most O157 strains show more expression of stx2 mRNA and protein than stx1. The low expression of stx2 gene implies that the stx expression may be influenced by environmental conditions of each strain.
- In conclusion, the present study demonstrates the diversity of virulence genes and serotypes in pathogenic E coli isolated from diarrhoea patients and the importance of stx2 gene in the infection of STEC O157. Thus, it can provide the epidemiological information regarding the trend of STEC and non-STEC infections in the general population and show the fundamental data in association of serotypes with virulence genes in diarrhoeagenic E coli strains from Korea.
Discussion
-
Acknowledgements
- This study was supported by a grant from Korean National Institute of Health, Seoul, Republic of Korea.
Acknowledgements
-
This is an Open Access article distributed under the terms of the Creative Commons Attribution Non-Commercial License (http://creativecommons.org/licenses/by-nc/3.0) which permits unrestricted non-commercial use, distribution, and reproduction in any medium, provided the original work is properly cited.
Article information
- 1. Clarke S.C.. Diarrhoeagenic Escherichia coli: an emerging problem? Diagn Microbiol Infect Dis 41:2001;93−99. PMID: 11750160.ArticlePubMed
- 2. Nataro J.P., Kaper J.B.. Diarrheagenic Escherichia coli. Clin Microbiol Rev 11:1998;142−201. PMID: 9457432.ArticlePubMedPMC
- 3. Cho S.H., Kim J.H., Kim J.C.. Surveillance of bacterial pathogens associated with acute diarrheal disease in the Republic of Korea during one year, 2003. J Microbiol 44:2006;327−335. PMID: 16820763.PubMed
- 4. Clarke S.C., Haigh R.D., Freestone P.P.E., Williams P.H.. Virulence of enteropathogenic Escherichia coli, a global pathogen. Clin Microbiol Rev 16:2003;365−378. PMID: 12857773.ArticlePubMedPMC
- 5. Crepin V.F., Junkal G., Frankel G.. Enteropathogenic and enterohemorrhagic Escherichia coli infections: translocation, translocation, translocation. Infect Immun 73:2005;2573−2585. PMID: 15845459.ArticlePubMedPMC
- 6. Osek J.. Development of a multiplex PCR approach for the identification of shiga toxin-producing Escherichia coli strains and their major virulence factor genes. J Appl Microbiol 95:2003;1217−1225. PMID: 14632994.ArticlePubMed
- 7. Sjöling Å, Wiklund G., Savarino S.J.. Comparative analyses of phenotypic and genotypic methods for detection of enterotoxigenic Escherichia coli toxins and colonization factors. J Clin Microbiol 45:2007;3295−3301. PMID: 17687011.ArticlePubMedPMC
- 8. Tarr P.I., Gordon C.A., Chandler W.L.. Shiga-toxin-producing Escherichia coli and haemolytic uraemic syndrome. Lancet 365:2005;1073−1086. PMID: 15781103.ArticlePubMed
- 9. Schmidt H., Zhang W.L., Kohler B.. Genetic diversity of intimin genes of attaching and effacing Escherichia coli strains. J Clin Microbiol 40:2002;4486−4492. PMID: 12454140.Article
- 10. Makino S.I., Tobe T., Asakura H.. Distribution of the secondary type III secretion system locus found in enterohemorrhagic Escherichia coli O157:H7 isolates among shiga toxin-producing E coli strains. J Clin Microbiol 41:2003;2341−2347. PMID: 12791847.ArticlePubMedPMC
- 11. Taylor K.A., O’Connell C.B., Luther P.W., Donnenberg M.S.. The EspB protein of enteropathogenic Escherichia coli is targeted to the cytoplasm of infected HeLa cells. Infect Immun 66:1998;5501−5507. PMID: 9784563.ArticlePubMedPMC
- 12. Dubreuil J.D., Menard L.P., Lussier J.G.. Expression, purification, and biochemical characterization of enteroaggregative Escherichia coli heat-stable enterotoxin 1. Prot Exp Puri 33:2004;223−231.Article
- 13. Nashikawa Y.Z., Hase A., Ogasawara J.. Diarrheagenic Escherichia coli isolated from stools of sporadic cases of diarrheal illness in Osaka city, Japan between 1997 and 2000: prevalence of enteroaggregateive E coli heat-stable enterotoxin 1 gene-possessing E. coli. Jpn J Infect Dis 55:2002;183−190. PMID: 12606826.PubMed
- 14. Griffiths M.W., Anand S.K.. Quorum sensing and expression of virulence in Escherichia coli O157:H7. Int J Food Microbiol 85:2003;1−9. PMID: 12810266.ArticlePubMed
- 15. Sperandio V., Torres A.G., Giron J.A., Kaper J.B.. Quorum sensing is a global regulatory mechanism in enterohemorrhagic Escherichia coli O157:H7. J Bacteriol 183:2001;5187−5197. PMID: 11489873.ArticlePubMedPMC
- 16. Walters M., Sperandio V.. Autoinducer 3 and epinephrine signaling in the knetics of locus of enterocyte effacement gene expression in enterohemorrhagic Escherichia coli. Infect Immun 74:2006;5445−5455. PMID: 16988219.ArticlePubMedPMC
- 17. Kim J.C., Yoon J.W., Kim J.B.. Omics-based analysis of the luxS mutation in a clinical isolate of Escherichia coli O157:H7 in Korea. J Microbiol Biotechnol 20:2010;415−419. PMID: 20208450.ArticlePubMed
- 18. Guinée P.A.M., Agterberg C.M., Jansen W.H.. Escherichia coli O antigen typing by means of a mechanized microtechnique. Appl Microbiol 24:1972;127−131. PMID: 4560465.ArticlePubMedPMC
- 19. O’brien A.D., Scott M.E., Melton-Celsa A.R.. Mutations in hns reduce the adherence of Shiga toxin-producing E coli O91:H21 strain B2F1 to human colonic epithelial cells and increase the production of hemolysin. Microb Pathog 34:2003;155−159. PMID: 12631477.ArticlePubMed
- 20. Boerlin P., Schmid H., Burnens A.P.. Verocytotoxin-producing Escherichia coli in patients with diarrhoea in Switzerland. Eur J Clin Microbiol Infect Dis 21:2002;810−813. PMID: 12461591.ArticlePubMed
- 21. Jenkins C., Chart H.. Serodiagnosis of infection with verocytotoxin-producing Escherichia coli. J Appl Microbiol 86:1999;569−575. PMID: 10212402.ArticlePubMed
References
| Pathogen (n) | Serogroups (n) |
|---|---|
| STEC (49) | O157 (13), O26 (10), O103 (6), O111 (4), O121 (4), O55 (3), O91(3), O117 (2), O2 (1), O21 (1), O104 (1), O119 (1) |
| EPEC (20) | OUT∗ (8), O26 (2), O1 (1), O11 (1), O110 (1), O119 (1), O139 (1), O142 (1), O152 (1), O159 (1), O35 (1), O63 (1) |
| EAEC (20) | OUT (3), O169 (2), O20 (2), O153 (2), O158 (1), O18 (1), O8 (1), O124 (1), O114 (1), O25 (1), O166 (1), O15 (1), O146 (1), O5 (1), O51 (1) |
| ETEC (20) | O6 (8), O20 (3), O8 (2), O25 (2), OUT (2) O9 (1), O15 (1), O102 (1) |
| EIEC (2) | OUT (2) |
Figure & Data
References
Citations

- Comparative Analysis of Human and Animal E. coli: Serotyping, Antimicrobial Resistance, and Virulence Gene Profiling
Mahmoud M. Bendary, Marwa I. Abdel-Hamid, Walaa A. Alshareef, Hanan M. Alshareef, Rasha A. Mosbah, Nasreen N. Omar, Mohammad M. Al-Sanea, Majid Alhomrani, Abdulhakeem S. Alamri, Walaa H. Moustafa
Antibiotics.2022; 11(5): 552. CrossRef - Antimicrobial peptide human β-defensin-2 improves in vitro cellular viability and reduces pro-inflammatory effects induced by enteroinvasive Escherichia coli in Caco-2 cells by inhibiting invasion and virulence factors’ expression
Alessandra Fusco, Vittoria Savio, Brunella Perfetto, Roberto Mattina, Giovanna Donnarumma
Frontiers in Cellular and Infection Microbiology.2022;[Epub] CrossRef - Distribution of Pathogenicity Island Markers and H-Antigen Types of Escherichia coli O25b/ST131 Isolates from Patients with Urinary Tract Infection in Iran
Masoumeh Rasoulinasab, Fereshteh Shahcheraghi, Mohammad Mehdi Feizabadi, Bahram Nikmanesh, Azade Hajihasani, Shahram Sabeti, Mohammad Mehdi Aslani
Microbial Drug Resistance.2021; 27(3): 369. CrossRef - Development and validation of a predictive model for pathogenic Escherichia coli in fresh‐cut produce
You Jin Kim, Ju Yeon Park, Soo Hwan Suh, Mi‐Gyeong Kim, Hyo‐Sun Kwak, Soon Han Kim, Eun Jeong Heo
Food Science & Nutrition.2021; 9(12): 6866. CrossRef - In vitro antibacterial activity of poly (amidoamine)-G7 dendrimer
Mitra Gholami, Rashin Mohammadi, Mohsen Arzanlou, Fakhraddin Akbari Dourbash, Ebrahim Kouhsari, Gharib Majidi, Seyed Mohsen Mohseni, Shahram Nazari
BMC Infectious Diseases.2017;[Epub] CrossRef - Occurrence of pathogenic Escherichia coli in commercially available fresh vegetable products in Korea
Hyun Jung Kim, Minseon Koo, A-Ram Jeong, Seung-Youb Baek, Joon-Il Cho, Soon-Ho Lee, In-Gyun Hwang
Journal of the Korean Society for Applied Biologic.2014; 57(3): 367. CrossRef


 PubReader
PubReader Cite
Cite
