Articles
- Page Path
- HOME > Osong Public Health Res Perspect > Volume 4(3); 2013 > Article
-
Original Article
Multiplex Real-time Polymerase Chain Reaction Assays for Simultaneous Detection ofVibrio cholerae ,Vibrio parahaemolyticus , andVibrio vulnificus - Jie Yeun Park, Semi Jeon, Jun Young Kim, Misun Park, Seonghan Kim
-
Osong Public Health and Research Perspectives 2013;4(3):133-139.
DOI: https://doi.org/10.1016/j.phrp.2013.04.004
Published online: April 30, 2013
Division of Enteric Bacterial Infections, Korea National Institute of Health, Osong, Korea
- ∗Corresponding author. kkingsh@chol.com
© 2013 Published by Elsevier B.V. on behalf of Korea Centers for Disease Control and Prevention.
This is an Open Access article distributed under the terms of the Creative Commons Attribution Non-Commercial License (http://creativecommons.org/licenses/by-nc/3.0) which permits unrestricted non-commercial use, distribution, and reproduction in any medium, provided the original work is properly cited.
Abstract
-
Objectives
- A multiplex real-time polymerase chain reaction (RT-PCR) method was developed for the identification of three Vibrio species: Vibrio cholerae, Vibrio parahaemolyticus, and Vibrio vulnificus.
-
Methods
- Specific primers and probes targeting the hlyA, tlh, and vvhA genes were selected and used for multiplex real-time PCR to confirm the identification of V. cholerae, V. parahaemolyticus, and V. vulnificus, respectively. This method was applied to screen Vibrio species from environmental samples and combining it with a culture-based method, its effectiveness was evaluated in comparison with culture-based methods alone.
-
Results
- Specific PCR fragments were obtained from isolates belonging to the target species, indicating a high specificity of this multiplex real-time PCR. No cross-reactivity with the assay was observed between the tested bacteria. The sensitivity of the multiplex real-time PCR was found to have a lower limit of 104 colony-forming units/reaction for all three Vibrio species. The combination strategy raised the isolation ratio of all three Vibrio species 1.26- to 2.75-fold.
-
Conclusion
- This assay provides a rapid, sensitive, and specific technique to detect these three Vibrio species in the environment.
- Vibrio is a genus of Gram-negative bacteria that possess a curved rod shape and naturally inhabits aquatic environments worldwide [1–5]. Within the genus Vibrio, several species are known to be important human pathogens [6–8]. Among these, Vibrio cholerae, Vibrio vulnificus, and Vibrio parahaemolyticus are the major pathogenic Vibrio species. V. cholerae and V. parahaemolyticus, contracted through consumption of contaminated seafood and seawater, can cause gastroenteritis whereas V. vulnificus can cause septicemia by exposure of an open wound to seawater or consumption of contaminated seafood [9–11]. In Korea especially, owing to the common practice of raw seafood consumption, gastroenteritis caused by infection with V. parahaemolyticus and septicemia caused by infection with V. vulnificus occur frequently.
- Because cases of infection by Vibrio spp. are commonly found in coastal areas, it is prudent to investigate the distribution of the pathogenic Vibrio spp. in the coastal environment of Korea. However, the isolation of bacteria using conventional microbiological method is a time-consuming and laborious process that presents potential chance for error resulting in nondetection of Vibrio spp. present in environmental samples.
- Real-time PCR is a rapid, sensitive, and highly specific technique. In many reports, real-time PCR has been used to detect various human and animal pathogens [12–19].
- In the present study, we developed a simple multiplex PCR method based on TaqMan real-time PCR to detect V. parahaemolyticus, V. vulnificus, and V. cholerae in a single PCR reaction. In this assay, we selected specific primers and probes targeting the hemolysin genes of three Vibrio spp. for their identification. The hlyA, tlh, and vvhA genes are species-specific markers for V. cholerae, V. parahaemolyticus, and V. vulnificus, respectively [20–23].
- This PCR method allowed for quick and easy isolation of three Vibrio species from environmental samples. Owing to its sensitivity, accuracy, and the potential increase in isolation rate in conjunction with other isolation methods, such as the use of chromogenic selective media, the method provides a rapid and effective detection tool for research and diagnostics.
Introduction
- 2.1 Bacterial strains and DNA extraction
- A total of 56 strains from the Korean Centers for Diseases Control and Prevention strain collection were used in this study (Table 1). These included five reference strains and 51 clinical and environmental strains. All strains were cultured on tryptic soy agar (Becton Dickinson and Company, Sparks, MD, USA) with overnight incubation at 37 °C.
- Genomic DNA extraction was performed by boiling. Briefly, one colony was suspended in 200 μL of sterile distilled water and boiled for 10 minutes to lyse the bacteria. After boiling, the suspension was centrifuged for 2 minutes at 14,000 rpm to sediment the cell debris. The supernatant was collected and used as a template for PCR.
- 2.2 Primer and TaqMan probe design
- Suitable primers and probes for the multiplex TaqMan real-time PCR (amplifying the hlyA, tlh, and vvhA genes of V. cholerae, V. parahaemolyticus, and V. vulnificus, respectively) were designed using Primer Express (Applied Biosystems, Foster City CA, USA) and were synthesized by Applied Biosystems (Table 2). Each set of primers was used to amplify its respective target gene from the purified genomic DNA of three Vibrio species reference strains (V. cholera ATCC 14033, V. parahaemolyticus ATCC 17802, and V. vulnificus ATCC 27562). All PCR products were verified by confirming their expected sizes via electrophoresis in a 3% Nusieve 3:1 agarose gel Lonza Group Ltd., Basel, Switzerland.
- 2.3 Multiplex TaqMan real-time PCR for detection of V. cholerae, V. parahaemolyticus, and V. vulnificus
- The real-time PCR reactions were performed on a Roche LightCycler 480 (Roche Diagnostics Ltd., Penzberg, Germany). Typical reactions contained 10 μL of 2× probes Master (Roche Diagnostics Ltd.), 90 nM each of the hlyA, tlh, and vvhA forward and reverse primers, 200 nM of the TaqMan MGB probes for hlyA, tlh, and vvhA (Table 2), and 1 uL of DNA template in a total volume of 20 μL. The cycling conditions were as follows: 2 minutes at 50 °C and 5 minutes at 95 °C, followed by 40 cycles, each consisting of 15 seconds at 95 °C and 1 minute at 60 °C. Data were analyzed using the LightCycler480 software (Roche Diagnostics Ltd.). PCR amplification was detected directly by monitoring the increase in fluorescence of each dye-labeled probe. Samples with cycle threshold (Ct) value above 35 were considered as negative.
- 2.4 Specificity and sensitivity of detection by multiplex TaqMan real-time PCR
- The specificity of each set of primers and probes for its respective target gene was determined by PCR amplification of the purified genomic DNA from 33 Vibrio spp. strains and 23 other bacterial strains.
- To identify the lower limit of detection and generate standard curves, 10-fold serial dilutions from 101 to 107 colony-forming units (CFU)/mL of each reference strain were used. Reference strains were cultured in tryptic soy broth media at 37 °C to yield 109 CFU/mL. Bacterial growth was monitored by observation of the absorbance at 600 nm using a spectrophotometer (GeneQuant pro; Amersham Pharmacia Biotech Inc., Cambridge, UK). Ten-fold serial dilutions of bacteria in physiological saline, from 101 to 107 CFU/mL, were used to generate templates for multiplex real-time PCR. To determine the number of cells in the samples, 100 μL of each 10-fold serial dilution was spread on a trypticase soy agar plate and incubated at 37 °C for 16 hours, and the grown bacterial colonies were counted. The standard curves were determined by plotting the Ct values against the log CFU/reaction. Amplification efficiency was calculated using the equation E=10[−1/slope] − 1.
- 2.5 Application of multiplex real-time PCR as a screening method
- Prior to the isolation of Vibrio species strains from the samples, real-time PCR was performed to detect the presence of V. cholerae, V. parahaemolyticus, and V. vulnificus in the samples. From May 2007 to December 2007, 2729 marine environmental samples (seawater, sediments, and plankton etc.) were collected from 11 Korean coastal areas. Ten milliliters or 10 g of the samples was added to 90 mL of alkaline peptone water (1% peptone, 1% NaCl, pH 8.4) and incubated 6–8 hours at 37 °C. After enrichment of the samples, a 1 mL portion of alkaline peptone water was removed, and the total DNA extracted by the boiling method previously described. The extracted DNA was used as a template for multiplex real-time PCR, as described above.
- A loopful of each enrichment culture was also streaked onto thiosulfate citrate bile salts sucrose (TCBS; Beckton Dickinson and Company) agar plates and incubated for 18–20 hours at 37 °C. After incubation, candidate colonies from the plate corresponding to the PCR positive culture were replica plated onto CHROMagar Vibrio (CV; CHROMagar, Paris, France) and TCBS agar plates. By comparing the growth on the TCBS and CV agar plates, a candidate species could be identified for each colony: a colony that was green on TCBS and blue on CV was thought to be V. vulnificus; a colony that was green on TCBS and mauve on CV was thought to be V. parahaemolyticus; and a colony that was yellow on TCBS and blue on CV was thought to be V. cholerae. The colonies were further identified biochemically using an API 20E identification kit (bioMérieux, Inc., Hazelwood, MO, USA).
Materials and Methods
- 3.1 Specificity and sensitivity of detection
- The primers and probes were specifically designed to identify the hemolysin genes present in V. cholerae, V. parahaemolyticus, and V. vulnificus: the hlyA, tlh, and vvhA genes, respectively. PCR amplification with these primers yielded amplicons of the expected molecular weights (57 bp for hlyA, 58 bp for tlh, and 79 bp for vvhA; Figure 1, Table 2).
- The specificity of the primers and probes against 33 Vibrio spp. strains and 23 other bacterial strains was tested by multiplex real-time PCR. Specificity was confirmed by PCR amplification of the hemolysin genes of V. cholerae, V. parahaemolyticus, and V. vulnificus, respectively, but no amplification was detected for any other Vibrio spp. or non-Vibrio spp. (Table 1).
- The aim of this study was to develop and evaluate a method for the simultaneous detection of V. cholerae, V. parahaemolyticus, and V. vulnificus using multiplex real-time PCR in a single sample. During development of the assay, no cross-reactivity between the tested bacteria was observed.
- To determine the detection limit of the multiplex real-time PCR and to establish a standard curve for quantification, DNA extracted from 10-fold serial dilutions of three Vibrio spp. at final concentration of 101 – 107 CFU/reaction was analyzed by real-time PCR. Standard curves were constructed for three Vibrio spp. using a single target DNA in the mixture of three primer sets and three fluorescent probes. When only one Vibrio species target was amplified by the multiplex real-time PCR assay, the detection limits were 103 CFU/reaction. The amplification efficiencies were 98% for V. cholerae, 97% for both V. parahaemolyticus, and V. vulnificus, resulting in highly accurate standard curves (Figure 2).
- Next, simultaneous detection of three Vibrio species was attempted using a mixture of DNA from the same three Vibrio species. As a consequence, the detection limit decreased by 10-fold (104 CFU/reaction) for all three Vibrio spp. However, the amplification efficiency was similar to the 98% achieved with a single target for all three Vibrio spp., indicating that this assay effectively quantified each target (Figure 3).
- 3.2 Application of real-time multiplex PCR as a screening method
- The multiplex real-time PCR assay developed in this study was used to screen samples from a pilot surveillance study of marine Vibrio in Korea. Of the 2729 marine environmental samples tested by the multiplex real-time PCR assay, 2085 (64.7%) were positive for the tlh gene, 621 (19.3%) were positive for the vvhA gene, and 330 (10.3%) were positive for the hlyA gene (Table 3).
- Isolation of the three Vibrio spp. was attempted from samples that had tested positive in the multiplex real-time PCR assay. The most prominent Vibrio species, V. parahaemolyticus, was isolated from 1501 (37%) of the 2769 samples, whereas V. cholerae and V. vulnificus were found in 228 (5.6%) and 180 (4.4%) of the samples, respectively. V. parahaemolyticus was the most frequently detected of the three Vibrio species using both culture-based methods and multiplex real-time PCR.
- The same surveillance project was performed in 2006, during the same seasonal period from May to December, prior to the development of multiplex PCR screening. In that study, V. parahaemolyticus was isolated from 1893 (29.3%) of the 5445 samples, followed by V. cholerae from 213 (3.3%), and V. vulnificus from 106 (1.6%; Table 3).
Results
- With climate change, the threat of huge outbreaks of disease caused by Vibrio species such as V. cholerae is increasing. Consequently, the importance of surveillance of pathogenic Vibrio species present in the environment is also growing [24]. However, isolation of specific Vibrio species from the environment is not easy due to the difficulty associated with discriminating between Vibrio species on selective media. Additionally, in culture-based methods, colony confirmation is usually carried out using a panel of biochemical tests. In a previous study, O’Hara et al reported that the API 20E identification kit possessed an accuracy >90% for V. parahaemolyticus identification, but it was able to identify only 50% and 60% of V. cholerae and V. vulnificus strains, respectively [25]. Because commercial bacterial identification systems are used for clinical isolates and a comprehensive evaluation of the ability of these systems to identify bacteria accurately from environmental samples has not been performed, commercial systems may misidentify Vibrio spp. as other bacterial species [26–28]. The phenotypic variability of environmental strains and the close relationship among Vibrio spp. account for the failure to identify Vibrio variants accurately. It has been reported that some V. cholerae strains are unable to ferment sucrose [29]. In 2004–2006, the V. cholerae variants isolated from four outbreaks in Taiwan were misidentified as V. mimicus and V. alginolyticus using an API 20E identification kit. However, these strains were correctly identified by several molecular techniques, including PCR [30].
- For these reasons, many molecular biological tools have been developed to detect Vibrio species more easily, and most of these are PCR-based methods [15,20,31]. These PCR methods are usually employed to detect only one species, and, even when multiplex PCR assays are used to detect multiple species, they are performed using conventional PCR methods, which have limited sensitivity and are time consuming [32,33].
- In the present study, we developed and tested a TaqMan probe-based multiplex real-time PCR assay to detect V. cholerae, V. vulnificus, and V. parahaemolyticus in environmental samples after enrichment. Because this multiplex real-time PCR utilizes three fluorescent probes simultaneously in a single PCR reaction, the speed at which three Vibrio spp. can be detected is greatly increased. This assay provides results within approximately 3 hours of enrichment because there is no need for postamplification analysis, such as agarose gel electrophoresis, for confirmation of real-time detection. In addition, the threshold cycles affords a further advantage of semiquantitation.
- We applied this multiplex real-time PCR method to screen marine environmental samples and performed culture-based isolation for the PCR-positive samples only. We compared the effectiveness of the combination of this multiplex real-time PCR and culture-based methods (samples collected in 2007) to culture-based methods alone (samples collected in 2006). The isolation ratios of V. cholerae, V. vulnificus, and V. parahaemolyticus were higher with the combination method than with the culture-based methods alone. The isolation ratio of V. vulnificus using combination methods was 2.75-fold higher than that of culture-based methods alone. The combination methods provided a higher isolation ratio than that of culture-based methods alone because we could neglect the samples that were PCR-negative and focus on which species to isolate from each of the PCR-positive samples.
- In conclusion, we developed a multiplex real-time PCR assay for the simultaneous detection of V. cholerae, V. parahaemolyticus, and V. vulnificus in a single reaction. The multiplex real-time PCR provides a rapid, sensitive, and highly specific means for the detection of three Vibrio spp. from environmental and clinical samples. This technique might be useful to detect the three pathogenic Vibrio spp. during mass outbreaks and sporadic cases of vibriosis, as well as in contaminated seafood and wastewater. It could facilitate monitoring of pathogenic Vibrio contamination, thereby improving hygiene.
Discussion
-
Acknowledgements
- This research was supported by a grant from the Korea Centers for Disease Control and Prevention.
Acknowledgments
-
This is an Open Access article distributed under the terms of the Creative Commons Attribution Non-Commercial License (http://creativecommons.org/licenses/by-nc/3.0) which permits unrestricted non-commercial use, distribution, and reproduction in any medium, provided the original work is properly cited.
Article information
- 1. Blackwell K.D., Oliver J.D.. The ecology of Vibrio vulnificus, Vibrio cholerae, and Vibrio parahaemolyticus in North Carolina estuaries. J Microbiol 46(2). 2008 Apr;146−153. PMID: 18545963.ArticlePubMed
- 2. Eiler A., Gonzalez-Rey C., Allen S., Bertilsson S.. Growth response of Vibrio cholerae and other Vibrio spp. to cyanobacterial dissolved organic matter and temperature in brackish water. FEMS Microbiol Ecol 60(3). 2007 Jun;411−418. PMID: 17386033.ArticlePubMed
- 3. Oliver J.D., Warner R.A., Cleland D.R.. Distribution of Vibrio vulnificus and other lactose-fermenting vibrios in the marine environment. Appl Environ Microbiol 45(3). 1983 Mar;985−998. PMID: 6847190.ArticlePubMed
- 4. Tamplin M., Rodrick G.E., Blake N.J., Cuba T.. Isolation and characterization of Vibrio vulnificus from two Florida estuaries. Appl Environ Microbiol 44(6). 1982 Dec;1466−1470. PMID: 7159088.ArticlePubMed
- 5. Wright A.C., Hill R.T., Johnson J.A., Roghman M.C., Colwell R.R., Morris J.G. Jr.. Distribution of Vibrio vulnificus in the Chesapeake Bay. Appl Environ Microbiol 62(2). 1996 Feb;717−724. PMID: 8593075.ArticlePubMed
- 6. Chakraborty S., Nair G.B., Shinoda S.. Pathogenic vibrios in the natural aquatic environment. Rev Environ Health 12(2). 1997 Apr–Jun;63−80. PMID: 9273923.ArticlePubMed
- 7. Hoge C.W., Watsky D., Peeler R.N., Libonati J.P., Israel E., Morris J.G. Jr.. Epidemiology and spectrum of Vibrio infections in a Chesapeake Bay community. J Infect Dis 160(6). 1989 Dec;985−993. PMID: 2584765.ArticlePubMed
- 8. Tarr C.L., Patel J.S., Puhr N.D., Sowers E.G., Bopp C.A., Strockbine N.A.. Identification of Vibrio isolates by a multiplex PCR assay and rpoB sequence determination. J Clin Microbiol 45(1). 2007 Jan;134−140. PMID: 17093013.ArticlePubMed
- 9. McLaughlin J.B., DePaola A., Bopp C.A.. Outbreak of Vibrio parahaemolyticus gastroenteritis associated with Alaskan oysters. N Engl J Med 353(14). 2005 Oct;1463−1470. PMID: 16207848.ArticlePubMed
- 10. Morris J.G. Jr.. Cholera and other types of vibriosis: a story of human pandemics and oysters on the half shell. Clin Infect Dis 37(2). 2003 Jul;272−280. PMID: 12856219.ArticlePubMed
- 11. Powell J.L.. Vibrio species. Clin Lab Med 19(3). 1999 Sep;537−552. vi. PMID: 10549425.ArticlePubMed
- 12. Gubala A.J.. Multiplex real-time PCR detection of Vibrio cholerae. J Microbiol Methods 65(2). 2006 May;278−293. PMID: 16153727.ArticlePubMed
- 13. Halse T.A., Musser K.A., Limberger R.J.. A multiplexed real-time PCR assay for rapid detection of Chlamydia trachomatis and identification of serovar L-2, the major cause of lymphogranuloma venereum in New York. Mol Cell Probes 20(5). 2006 Oct;290−297. PMID: 16644182.ArticlePubMed
- 14. McDonough E.A., Barrozo C.P., Russell K.L., Metzgar D.. A multiplex PCR for detection of Mycoplasma pneumoniae, Chlamydophila pneumoniae, Legionella pneumophila, and Bordetella pertussis in clinical specimens. Mol Cell Probes 19(5). 2005 Oct;314−322. PMID: 16024220.ArticlePubMed
- 15. Panicker G., Bej A.K.. Real-time PCR detection of Vibrio vulnificus in oysters: comparison of oligonucleotide primers and probes targeting vvhA. Appl Environ Microbiol 71(10). 2005 Oct;5702−5709. PMID: 16204478.ArticlePubMed
- 16. Qvarnstrom Y., Visvesvara G.S., Sriram R., da Silva A.J.. Multiplex real-time PCR assay for simultaneous detection of Acanthamoeba spp., Balamuthia mandrillaris, and Naegleria fowleri. J Clin Microbiol 44(10). 2006 Oct;3589−3595. PMID: 17021087.ArticlePubMed
- 17. Takahashi H., Hara-Kudo Y., Miyasaka J., Kumagai S., Konuma H.. Development of a quantitative real-time polymerase chain reaction targeted to the toxR for detection of Vibrio vulnificus. J Microbiol Methods 61(1). 2005 Apr;77−85. PMID: 15676198.ArticlePubMed
- 18. Ward L.N., Bej A.K.. Detection of Vibrio parahaemolyticus in shellfish by use of multiplexed real-time PCR with TaqMan fluorescent probes. Appl Environ Microbiol 72(3). 2006 Mar;2031−2042. PMID: 16517652.ArticlePubMed
- 19. Wolf S., Williamson W.M., Hewitt J.. Sensitive multiplex real-time reverse transcription-PCR assay for the detection of human and animal noroviruses in clinical and environmental samples. Appl Environ Microbiol 73(17). 2007 Sep;5464−5470. PMID: 17616614.ArticlePubMed
- 20. Lyon W.J.. TaqMan PCR for detection of Vibrio cholerae O1, O139, non-O1, and non-O139 in pure cultures, raw oysters, and synthetic seawater. Appl Environ Microbiol 67(10). 2001 Oct;4685−4693. PMID: 11571173.ArticlePubMed
- 21. Nordstrom J.L., Vickery M.C., Blackstone G.M., Murray S.L., DePaola A.. Development of a multiplex real-time PCR assay with an internal amplification control for the detection of total and pathogenic Vibrio parahaemolyticus bacteria in oysters. Appl Environ Microbiol 73(18). 2007 Sep;5840−5847. PMID: 17644647.ArticlePubMed
- 22. Wright A.C., Morris J.G. Jr., Maneval D.R. Jr., Richardson K., Kaper J.B.. Cloning of the cytotoxin-hemolysin gene of Vibrio vulnificus. Infect Immun 50(3). 1985 Dec;922−924. PMID: 4066036.ArticlePubMed
- 23. Wu Z.H., Lou Y.L., Lu Y.Y., Yan J.. Development of quantitative real-time polymerase chain reaction for the detection of Vibrio vulnificus based on hemolysin (vvhA) coding system. Biomed Environ Sci 21(4). 2008 Aug;296−301. PMID: 18837292.ArticlePubMed
- 24. Emch M., Feldacker C., Islam M.S., Ali M.. Seasonality of cholera from 1974 to 2005: a review of global patterns. Int J Health Geogr 7:2008 Jun;31PMID: 18570659.ArticlePubMed
- 25. O'Hara C.M., Sowers E.G., Bopp C.A., Duda S.B., Strockbine N.A.. Accuracy of six commercially available systems for identification of members of the family Vibrionaceae. J Clin Microbiol 41(12). 2003 Dec;5654−5659. PMID: 14662957.ArticlePubMed
- 26. Croci L., Suffredini E., Cozzi L.. Comparison of different biochemical and molecular methods for the identification of Vibrio parahaemolyticus. J Appl Microbiol 102(1). 2007 Jan;229−237. PMID: 17184339.ArticlePubMed
- 27. MacDonell M.T., Singleton F.L., Hood M.A.. Diluent composition for use of API 20E in characterizing marine and estuarine bacteria. Appl Environ Microbiol 44(2). 1982 Aug;423−427. PMID: 7125655.ArticlePubMed
- 28. Martinez-Urtaza J., Lozano-Leon A., Viña-Feas A., de Novoa J., Garcia-Martin O.. Differences in the API 20E biochemical patterns of clinical and environmental Vibrio parahaemolyticus isolates. FEMS Microbiol Lett 255(1). 2006 Feb;75−81. PMID: 16436064.ArticlePubMed
- 29. Ansaruzzaman M., Rahman M., Kibriya A.K., Bhuiyan N.A., Islam M.S., Albert M.J.. Isolation of sucrose late-fermenting and nonfermenting variants of Vibrio cholerae O139 Bengal: implications for diagnosis of cholera. J Clin Microbiol 33(5). 1995 May;1339−1340. PMID: 7615751.ArticlePubMed
- 30. Wei S.W., Chern L.L., Wu Y.C., Wang Y.L., Lin C.M., Chiou C.S.. Foodborne disease outbreaks caused by sucrose-nonfermenting and beta-galactosidase-deficient variants of Vibrio cholerae. Int J Food Microbiol 122(1–2). 2008 Feb;148−155. PMID: 18164089.ArticlePubMed
- 31. Panicker G., Myers M.L., Bej A.K.. Rapid detection of Vibrio vulnificus in shellfish and Gulf of Mexico water by real-time PCR. Appl Environ Microbiol 70(1). 2004 Jan;498−507. PMID: 14711681.ArticlePubMed
- 32. Aridgides L.J., Doblin M.A., Berke T., Dobbs F.C., Matson D.O., Drake L.A.. Multiplex PCR allows simultaneous detection of pathogens in ships' ballast water. Mar Pollut Bull 48(11–12). 2004 Jun;1096−1101. PMID: 15172815.ArticlePubMed
- 33. Bauer A., Rørvik L.M.. A novel multiplex PCR for the identification of Vibrio parahaemolyticus, Vibrio cholerae and Vibrio vulnificus. Lett Appl Microbiol 45(4). 2007 Oct;371−375. PMID: 17897378.ArticlePubMed
References
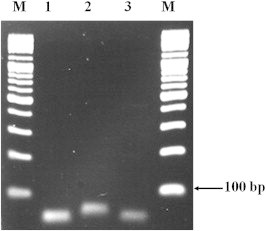
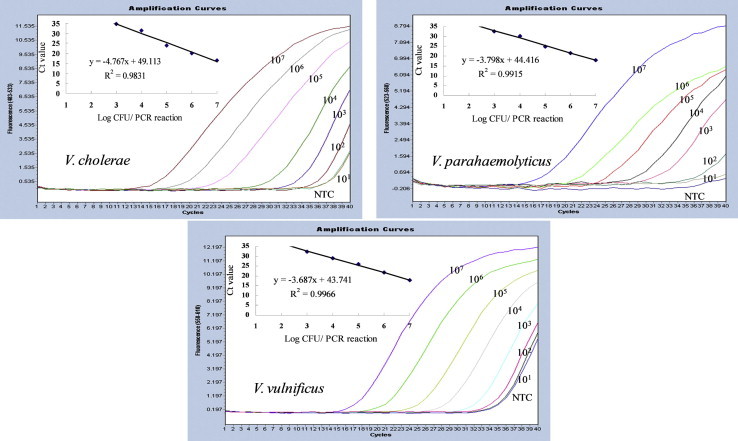
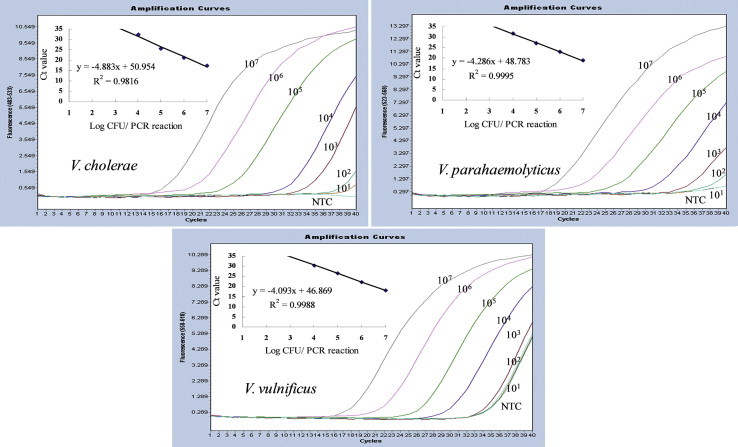
Figure & Data
References
Citations

- Vibrio cholerae and Salmonella Typhi culture-based wastewater or non-sewered sanitation surveillance in a resource-limited region
Petros Chigwechokha, Ruth Lusungu Nyirenda, Davie Dalitsani, Ranken Lorvin Namaumbo, Yohanny Kazembe, Ted Smith, Rochelle H. Holm
Journal of Exposure Science & Environmental Epidem.2024;[Epub] CrossRef - Incidence, virulence genes and antimicrobial resistance of Vibrio parahaemolyticus isolated from seafood
Deyan Stratev, Rumyana Fasulkova, Gergana Krumova-Valcheva
Microbial Pathogenesis.2023; 177: 106050. CrossRef - Exploring the Effect of Functional Diets Containing Phytobiotic Compounds in Whiteleg Shrimp Health: Resistance to Acute Hepatopancreatic Necrotic Disease Caused by Vibrio parahaemolyticus
Carla Hernández-Cabanyero, Esther Carrascosa, Silvia Jiménez, Belén Fouz
Animals.2023; 13(8): 1354. CrossRef - Multiplex PCR-Lateral Flow Dipstick Method for Detection of Thermostable Direct Hemolysin (TDH) Producing V. parahaemolyticus
Jirakrit Saetang, Phutthipong Sukkapat, Suriya Palamae, Prashant Singh, Deep Nithun Senathipathi, Jirayu Buatong, Soottawat Benjakul
Biosensors.2023; 13(7): 698. CrossRef - Current trends in polymerase chain reaction based detection of three major human pathogenic vibrios
Sharmin Quazi Bonny, M. A. Motalib Hossain, Syed Muhammad Kamal Uddin, Thiruchelvi Pulingam, Suresh Sagadevan, Mohd Rafie Johan
Critical Reviews in Food Science and Nutrition.2022; 62(5): 1317. CrossRef - Highly lethal Vibrio parahaemolyticus strains cause acute mortality in Penaeus vannamei post-larvae
Feng Yang, Limei Xu, Wanzhen Huang, Fang Li
Aquaculture.2022; 548: 737605. CrossRef - Hepatopancreatic transcriptome analysis and humoral immune factor assays in red claw crayfish (Cherax quadricarinatus) provide insight into innate immunomodulation under Vibrio parahaemolyticus infection
Duanduan Chen, Leifeng Guo, Cao Yi, Shouquan Wang, Yuanyuan Ru, Hui Wang
Ecotoxicology and Environmental Safety.2021; 217: 112266. CrossRef - Molecular mechanisms of Vibrio parahaemolyticus pathogenesis
Lingzhi Li, Hongmei Meng, Dan Gu, Yang Li, Mengdie Jia
Microbiological Research.2019; 222: 43. CrossRef - Application of digital PCR and next generation sequencing in the etiology investigation of a foodborne disease outbreak caused by Vibrio parahaemolyticus
Ying Li, Shuang Zhang, Jie Li, Meiling Chen, Mu He, Yuanyuan Wang, Yanchun Zhang, Hongbo Jing, Hongmei Ma, Yindong Li, Lin Zhao, Hongqun Zhao, Biao Kan, Bo Pang
Food Microbiology.2019; 84: 103233. CrossRef - Cholera Outbreak due to Raw Seafood Consumption in South Korea, 2016
Jeong Hyun Kim, Jin Lee, Sahyun Hong, Sangwon Lee, Hae-young Na, Young-Il Jeong, Eun Jin Choi, Junyoung Kim, Hyo Sun Kawk, Enhi Cho
The American Journal of Tropical Medicine and Hygi.2018; 99(1): 168. CrossRef - Detection and differentiation of Vibrio parahaemolyticus by multiplexed real-time PCR
Deshun Xu, Lei Ji, Xiaofang Wu, Wei Yan, Liping Chen
Canadian Journal of Microbiology.2018; 64(11): 809. CrossRef - Development of a rapid PCR protocol to detect Vibrio parahaemolyticus in clams
Sara Federici, Diana I. Serrazanetti, M. Elisabetta Guerzoni, Raffaella Campana, Eleonora Ciandrini, Wally Baffone, Andrea Gianotti
Journal of Food Science and Technology.2018; 55(2): 749. CrossRef - First Case of Necrotizing Fasciitis Caused by Skermanella aerolata Infection Mimicking Vibrio Sepsis
Sang Taek Heo, Ki Tae Kwon, Jeong Rae Yoo, Ji Young Choi, Keun Hwa Lee, Kwan Soo Ko
Annals of Laboratory Medicine.2018; 38(6): 604. CrossRef - Vibrio cholerae O1 with Reduced Susceptibility to Ciprofloxacin and Azithromycin Isolated from a Rural Coastal Area of Bangladesh
Shah M. Rashed, Nur A. Hasan, Munirul Alam, Abdus Sadique, Marzia Sultana, Md. Mozammel Hoq, R. Bradley Sack, Rita R. Colwell, Anwar Huq
Frontiers in Microbiology.2017;[Epub] CrossRef - A label-free multi-functionalized graphene oxide based electrochemiluminscence immunosensor for ultrasensitive and rapid detection of Vibrio parahaemolyticus in seawater and seafood
Yuhong Sha, Xuan Zhang, Wenrou Li, Wei Wu, Sui Wang, Zhiyong Guo, Jun Zhou, Xiurong Su
Talanta.2016; 147: 220. CrossRef - Toxigenic Vibrio cholerae O1 in vegetables and fish raised in wastewater irrigated fields and stabilization ponds during a non-cholera outbreak period in Morogoro, Tanzania: an environmental health study
Yaovi M. G. Hounmanou, Robinson H. Mdegela, Tamègnon V. Dougnon, Ofred J. Mhongole, Edward S. Mayila, Joseph Malakalinga, George Makingi, Anders Dalsgaard
BMC Research Notes.2016;[Epub] CrossRef - Isolation of Vibrio vulnificus Biotype I from Disease Outbreaks on Cultured Tiger Grouper Epinephelus fuscoguttatus Forsskal, 1775
Jumroensri Thawonsuwan, Jiraporn Kasornchandra, Patcharee Soonsan, Chantana Keawtapee
Fish Pathology.2016; 51(Special-is): S39. CrossRef - Detection of Vibrio cholerae by isothermal cross-priming amplification combined with nucleic acid detection strip analysis
Xia Zhang, Xin-Jun Du, Chun Guan, Ping Li, Wen-Jie Zheng, Shuo Wang
Molecular and Cellular Probes.2015; 29(4): 208. CrossRef - lolB gene, a valid alternative for qPCR detection of Vibrio cholerae in food and environmental samples
Alejandro Garrido-Maestu, María-José Chapela, Juan M. Vieites, Ana G. Cabado
Food Microbiology.2015; 46: 535. CrossRef


 PubReader
PubReader Cite
Cite
